Scientists at Tokyo Tech have produced computational DNA droplets that can recognize specific patterns in tumor biomarker microRNA sequences by merging the technologies of DNA droplets and DNA computing. When integrated into biomolecular robots and artificial cells, the technology will have potential applications in cancer diagnosis, drug resistance analysis, and residual tumor detection.
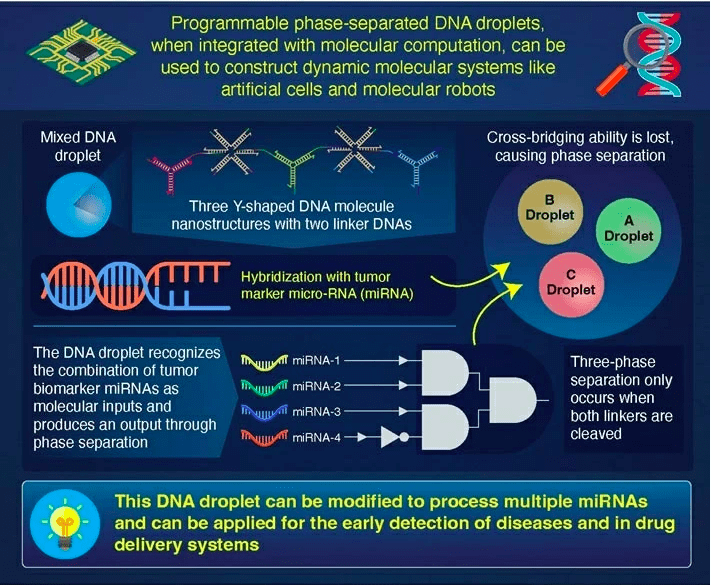
DNA droplets are generated by the liquid-liquid phase separation of macromolecules and play an essential role in regulating biological processes. This mechanism can be utilized to build self-assembled dynamic molecular systems, such as artificial cells and molecular robots.
The production of aqueous droplets in macromolecules by liquid-liquid phase separation (or coacervation) is a hot topic in biosciences studies. DNA is particularly interesting among the different macromolecules that form droplets because it is predictable and programmable, both of which are desirable properties in nanotechnology. The programmability of DNA has recently been used to build and govern DNA droplets created by the coacervation of sequence-designed DNAs.
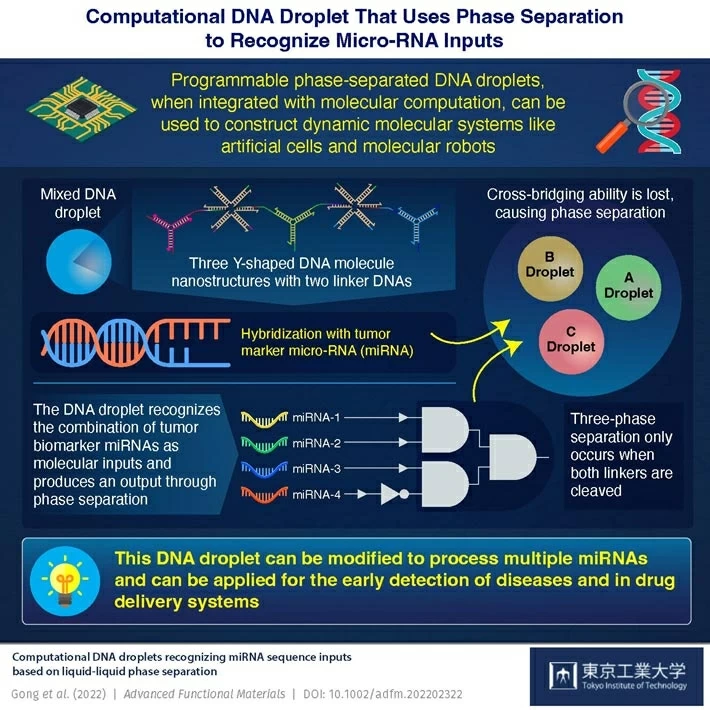
Prof. Masahiro Takinoue’s team at Tokyo University of Technology (Tokyo Tech) has developed a computational DNA droplet that can distinguish certain combinations of chemically generated microRNAs (miRNAs) that act as tumor indicators. Through physical DNA droplet phase separation, the droplets can produce a DNA logic computation output using these miRNAs as a molecular input.
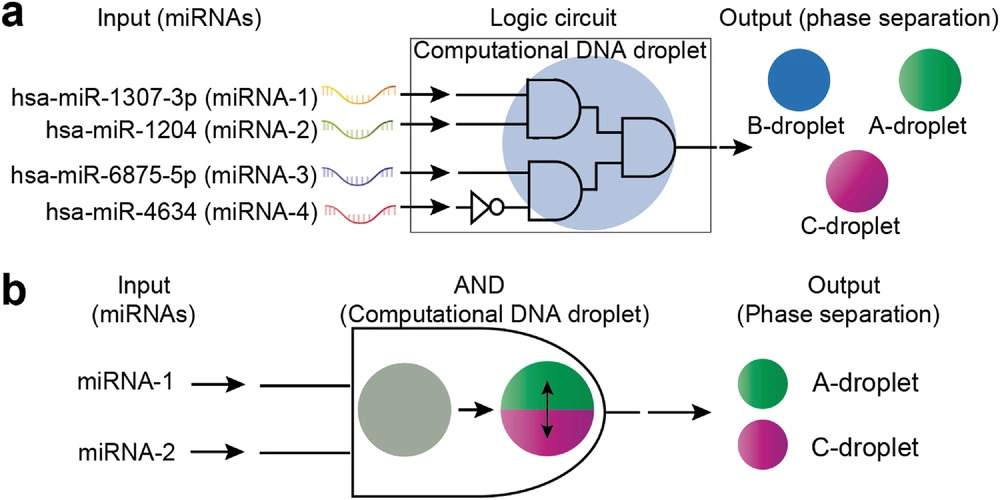
Image Source: Computational DNA Droplets Recognizing miRNA Sequence Inputs Based on Liquid–Liquid Phase Separation.
According to Prof. Takinoue, such research is necessary as DNA droplets have been used in cell-inspired microcompartments before. There is no literature on DNA droplet integration with molecular computing, despite the fact that biological systems regulate their functioning by integrating biosensing with molecular logical processing. The findings of the research were published in Advanced Functional Materials.
A number of experiments were performed to create this DNA droplet. To construct A, B, and C DNA droplets, scientists first created three types of Y-shaped DNA nanostructures termed Y-motifs A, B, and C, each having three sticky ends. Similar droplets tend to stick together naturally, whereas different droplets require the use of a particular “linker” molecule. Combining the A droplet with the B and C droplets employed linker molecules, which were referred to as AB and AC linkers, respectively.
The researchers tested the “AND” action in the AB droplet mixture by introducing two input DNAs in the first experiment. The presence of input is recorded as 1 in this procedure, while its absence is recorded as 0. The phase separation of the AB droplet mixture occurred only at (1,1), that is, when both input DNAs are present, indicating that the AND operation was successfully applied. Following this research, the researchers chose to add breast cancer tumor markers, miRNA-1 and miRNA-2, to the AC droplet combination as AND inputs. Because the AND operation was effective, the computational DNA droplet was able to identify the miRNAs.
Next, the researchers demonstrated simultaneous AND and NOT operations in an AB mixture with miRNA-3 and miRNA-4 breast cancer biomarkers in subsequent tests. Finally, they made an ABC droplet combination and added each of the four breast cancer indicators to it. The phase separation in an ABC droplet was determined by the linker cleavage, which resulted in either a two-phase or three-phase separation.
The researchers demonstrated the ability to identify a set of recognized cancer biomarkers or markers for three diseases simultaneously thanks to this property of the ABC droplet. Prof. Takinoue, also the corresponding author of the paper, believes that computational DNA droplets have enormous promise.
If a DNA droplet can be developed which can integrate and process multiple inputs and outputs, we can use it in early disease detection as well as drug delivery systems. Our current study also acts as a steppingstone for research in developing intelligent artificial cells and molecular robots.
Prof. Masahiro Takinoue
Story Source: Gong, J., Tsumura, N., Sato, Y., & Takinoue, M. (2022). Computational DNA Droplets Recognizing miRNA Sequence Inputs Based on Liquid–Liquid Phase Separation. Advanced Functional Materials, 2202322.
https://www.titech.ac.jp/english/news/2022/064204
Dr. Tamanna Anwar is a Scientist and Co-founder of the Centre of Bioinformatics Research and Technology (CBIRT). She is a passionate bioinformatics scientist and a visionary entrepreneur. Dr. Tamanna has worked as a Young Scientist at Jawaharlal Nehru University, New Delhi. She has also worked as a Postdoctoral Fellow at the University of Saskatchewan, Canada. She has several scientific research publications in high-impact research journals. Her latest endeavor is the development of a platform that acts as a one-stop solution for all bioinformatics related information as well as developing a bioinformatics news portal to report cutting-edge bioinformatics breakthroughs.