A new study reveals how antibodies choose which antigens to bind to while navigating the surface of pathogens such as coronaviruses. Scientists from KTH Royal Institute of Technology and Karolinska Institutet have developed a model that implies pathogen hunters move like a child playing in a stream with stepping stones.
The study displays antibody–antigen interactions as a computational system in which antigen geometry constrains and potentially controls the antibody movement.
According to Ian Hoffecker, a KTH researcher, the model introduces new ways to think about the virus and immune system evolution. The new study yields insights that could be valuable in vaccine engineering.
Antibodies are frequently shown as Y-shaped proteins. But recent research suggests that a more precise way to view them is to turn the picture upside down and consider antibodies as walking stick figures, stepping on antigens. Hoffecker says the two distinctive “Y” branches serve as “legs.”
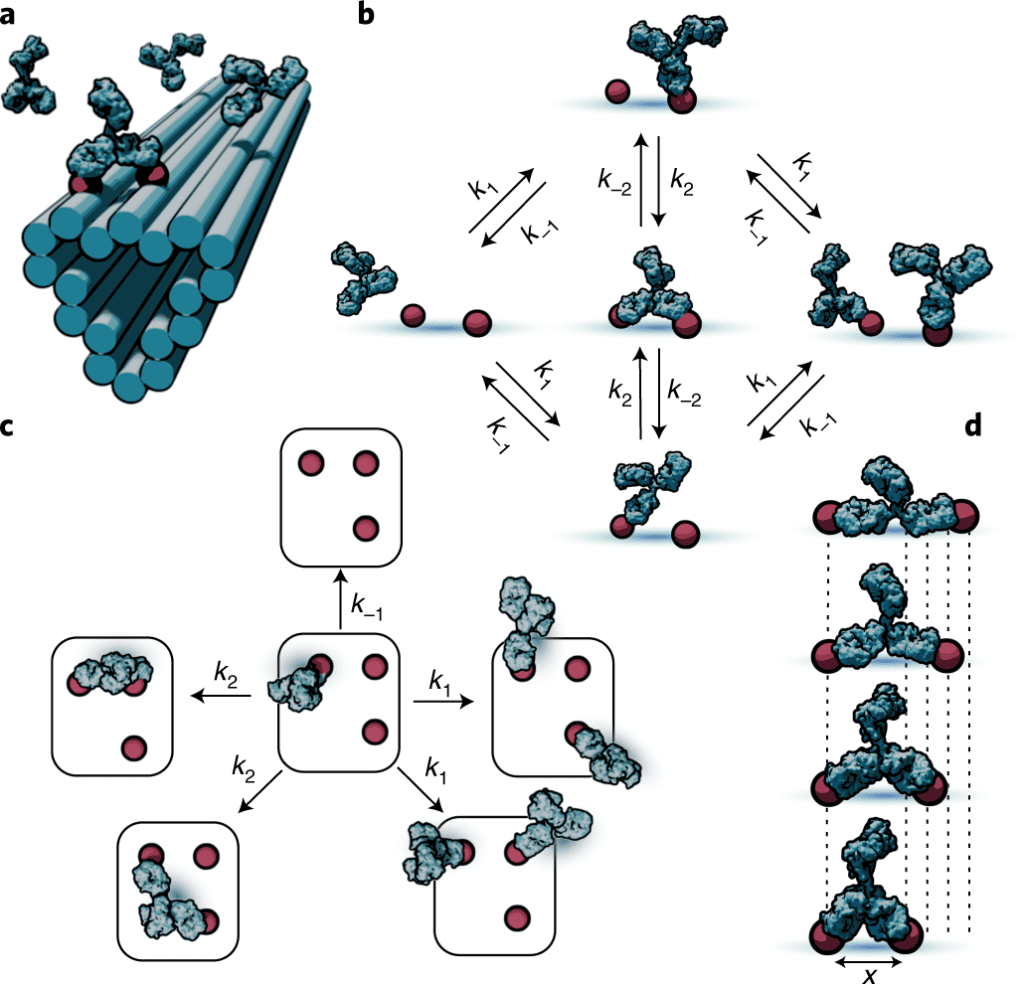
Image Source: Stochastic modeling of antibody binding predicts programmable migration on antigen patterns.
“These antibodies are built for walking,” he explains, paraphrasing Nancy Sinatra’s 1966 hit recording.
These stalking pathogen hunters identify their target by planting their “feet” on antigens, which are tiny molecules distributed on the surfaces of viruses like stepping stones in varied patterns. They rely on a process known as multivalence, which allows them to bind to their targets as strongly as possible by establishing a footing with both “Y” branches, generally on two different antigens. Once established, the antibodies interact with other signaling proteins to neutralize or kill the pathogen.
According to Hoffecker, the researchers used a nano-fabricated model of a pathogen’s antigen pattern to see how pathogen surfaces influence this behavior. He adds, “What if the antigens are relatively near together or far apart?” Do the molecules of antibodies stretch out or compress?”
To find out, Björn Högberg of Karolinska Institutet’s Division of Biomaterials Research says the team used a method known as DNA origami, in which DNA self-assembles into nanostructures with programmable geometry, allowing them to manipulate the distance between antigens, to simulate a pathogen and antigen scenario.
This tool has enabled how to investigate the distance between two antigens that impacts the binding strength. This data was plugged into a model that allows posing intriguing questions about how antibodies behave in more complex situations while staying true to reality.
According to Hoffecker, the model demonstrates that antibodies behave similarly to another well-known bipedal organism, humans.
“The process could be likened to a child playing on a river laden with stepping stones just large enough to accommodate a single foot,” Hoffecker says. “So to stand in place, the child would have to straddle two rocks or else balance on one foot.”
The model’s antibodies seemed to prefer antigens closer together and easier to stand on. Antigens that are too far apart have a statistical tendency to move to a region where they are closer together, according to Hoffecker.
Such findings raise the question of whether antibodies’ flexibility and structure are modified by their antagonists, the pathogen. “We are asking the question, is this relevant to evolution, or co-evolution, where you have this constant arms race between the immune system and pathogens and this control system that basically says how antibodies move and where they go?” Hoffecker says.
According to Hoffecker, the next step is to investigate how this antibody trait expresses itself in natural systems and incorporate these discoveries into rationally designed vaccinations that account for the antigen spatial organizing element.
Story Source: Hoffecker, I.T., Shaw, A., Sorokina, V. et al. Stochastic modeling of antibody binding predicts programmable migration on antigen patterns. Nat Comput Sci 2, 179–192 (2022). https://doi.org/10.1038/s43588-022-00218-z https://www.kth.se/en/aktuellt/nyheter/how-antibodies-navigate-pathogen-surfaces-like-a-child-at-play-1.1155230
Dr. Tamanna Anwar is a Scientist and Co-founder of the Centre of Bioinformatics Research and Technology (CBIRT). She is a passionate bioinformatics scientist and a visionary entrepreneur. Dr. Tamanna has worked as a Young Scientist at Jawaharlal Nehru University, New Delhi. She has also worked as a Postdoctoral Fellow at the University of Saskatchewan, Canada. She has several scientific research publications in high-impact research journals. Her latest endeavor is the development of a platform that acts as a one-stop solution for all bioinformatics related information as well as developing a bioinformatics news portal to report cutting-edge bioinformatics breakthroughs.
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar







