Machine learning algorithms can provide precise forecasting of virus propagation during pandemics. This method eliminates human bias and assumptions, allowing for a more accurate prediction of pandemic evolution.
Researchers from KAUST developed a dynamic machine learning model with an efficient forecasting approach that can predict confirmed and recovered COVID-19 times series data with high accuracy.
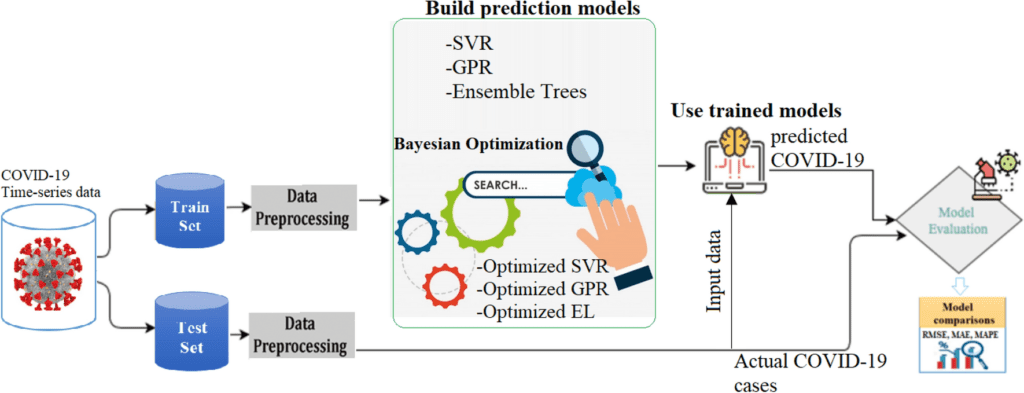
Image Source: A proficient approach to forecast COVID-19 spread via optimized dynamic machine learning models.
According to a new KAUST study, machine learning approaches can achieve an assumption-free analysis of epidemic case data with amazingly good prediction accuracy and the flexibility to incorporate new data dynamically. Yasminah Alali, an intern in KAUST’s 2021 Saudi Summer Internship (SSI) program, developed a proof of concept that reveals a possible alternative to traditional parameter-driven mechanistic models by removing human bias and assumptions from analysis, revealing the underlying story of the data.
Using publicly released COVID-19 incidence and recovery data from India and Brazil, Alali leveraged her experience working with artificial intelligence models to design a framework to fit the characteristics and time-evolving nature of epidemic data in collaboration with KAUST’s Ying Sun and Fouzi Harrou.
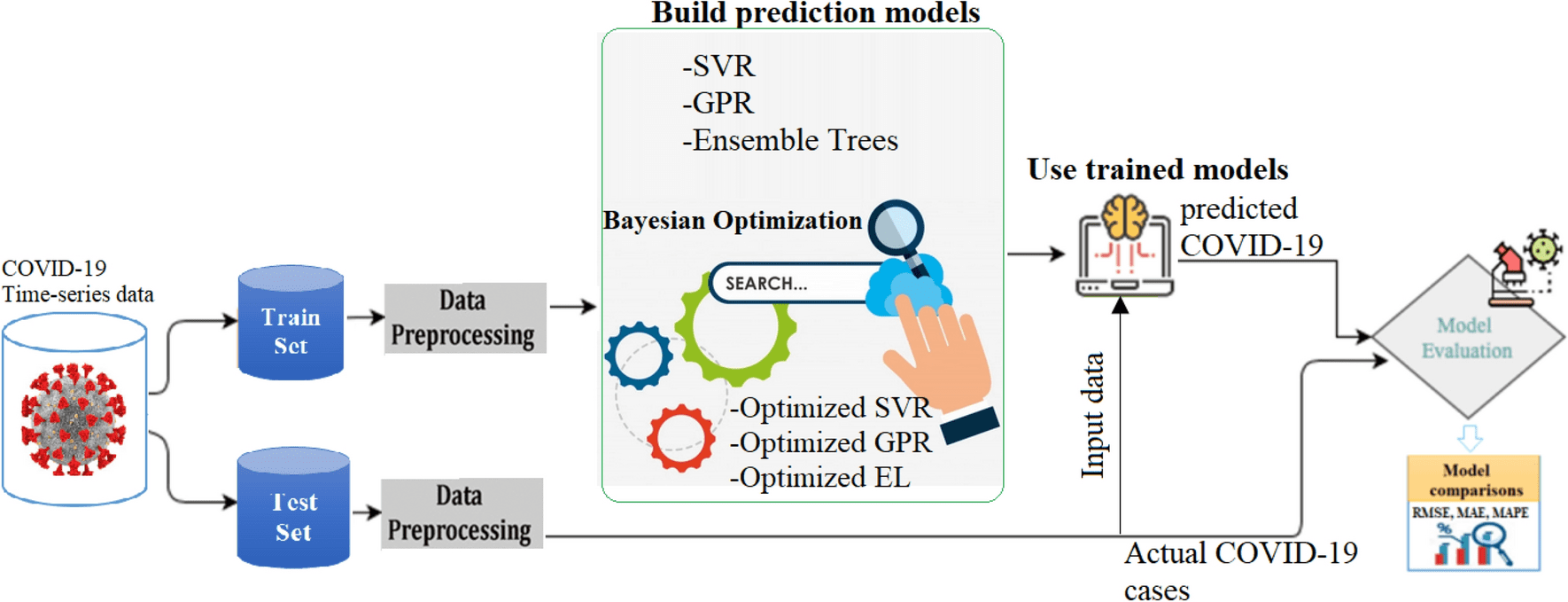
To create an effective Gaussian process regression (GPR) based model for forecasting recovered and confirmed COVID-19 cases in two significantly impacted countries, India and Brazil, the researchers first used Bayesian optimization to modify the Gaussian process regression (GPR) hyperparameters.
However, the time dependency in the COVID-19 data series is ignored by machine learning models. By integrating lagged measurements in the construction of the researched machine learning models, dynamic information has been taken into account to ameliorate this constraint. Using the Random Forest approach, the researchers also evaluated the contribution of the integrated features to the COVID-19 prediction.
Accurate case forecasting is critical for mitigating and slowing the spread during a pandemic. Various mathematical and time-series models have been created to enhance case spread forecasts, but these rely on a mechanical knowledge of how contagion spreads and the efficacy of mitigation measures like masks and isolation. As the understanding of a particular contagion grows, such methods become more accurate, yet this might lead to erroneous assumptions that unwittingly influence the modeling results’ accuracy.
The team had to come up with a means to dynamically include new data at different times in the learning process by “lagging” the data inputs because machine learning models can’t capture the time dependence of a data series. They also used a Bayesian optimization method to fine-tune the derived distributions for better precision. The end product is an integrated dynamic ML system that outperformed real-world data.
According to Harrou, machine learning algorithms are used in this study because they can extract considerable information from data flexibly and without making assumptions about the underlying data distribution. GPR is particularly appealing for processing many types of data with diverse Gaussian or nonGaussian distributions, and the integration of lagged data improves prediction quality significantly.
Story Source: Alali, Y., Harrou, F. & Sun, Y. A proficient approach to forecast COVID-19 spread via optimized dynamic machine learning models. Sci Rep 12, 2467 (2022). https://doi.org/10.1038/s41598-022-06218-3
https://discovery.kaust.edu.sa/en/article/1235/can-machine-learning-help-predict-disease-spread%3F
Dr. Tamanna Anwar is a Scientist and Co-founder of the Centre of Bioinformatics Research and Technology (CBIRT). She is a passionate bioinformatics scientist and a visionary entrepreneur. Dr. Tamanna has worked as a Young Scientist at Jawaharlal Nehru University, New Delhi. She has also worked as a Postdoctoral Fellow at the University of Saskatchewan, Canada. She has several scientific research publications in high-impact research journals. Her latest endeavor is the development of a platform that acts as a one-stop solution for all bioinformatics related information as well as developing a bioinformatics news portal to report cutting-edge bioinformatics breakthroughs.
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar