OpenProtein.AI is an open-access platform founded by MIT alumni Tristan Bepler and Tim Lu, which provides open-source AI models and a no-code interface for protein engineering. With a motivation to democratize access to advanced protein language models like PoET and PoET-2, they aim to enable biologists without coding expertise to design and analyze proteins by adding user-friendly tools that broaden the reach of biotech innovation.
In our earlier blogs, we’ve discussed various AI-integrated models for different biological tasks, and indeed, AI has proven itself well enough in speeding up drug development and deepening our understanding of disease mechanisms.
However, most scientists in the biotechnology and pharmaceutical industries lack expertise in machine learning techniques. These scientists find it difficult to apply such AI technologies because they have no proper training in them.
The researchers behind OpenProtein.AI argue that AI in biology isn’t just about technology, but also accessibility. Their platform is designed to put the most advanced protein models into the hands of every scientist, not just machine learning specialists, so that drug discovery can move faster.
AI as a Biologist: The Motivation and Backstory Behind OpenProtein.AI
The cofounder of OpenProtein.AI, Tristan Bepler, emphasizes in one of his recent LinkedIn posts that a powerful AI model reflects a deeper grasp of evolutionary rules that make up the proteins. He often highlights that instead of scientists manually articulating, the models uncover them directly from the sequence data, shifting them from just being a tool to a ‘knowledge engine’. He stresses that bridging deep AI principles and practical workflows is the way forward.
Bepler began his journey at MIT by analyzing evolutionary data of billions of proteins, before Google’s AlphaFold revolutionized protein structure prediction. He was among the first to treat proteins as a ‘language’ where sequences encode rules of structure and function.
As we know by now, the classical approach for designing a protein is to obtain the sequence, develop the structure, and map the functions. But we don’t fully understand how structure maps to function, and predicting structure is also computationally expensive.
Bepler’s idea is to use foundation models to bypass the intermediate ‘structure’ step and directly move from sequence to the function of proteins. This could allow scientists to design proteins with desired functions without solving their 3D structures.
Around the mid-2010s, the idea of integrating AI with biology was just gaining traction. This was when both the founders realized that the most advanced AI tools were inaccessible to most biologists because they required coding, GPUs, and specialized knowledge.
How OpenProtein.AI Serves As a Flexible, Widely Usable Platform
OpenProtein.AI turns protein language models like PoET (Protein Evolutionary Transformer) into practical, everyday tools for biologists. PoET was trained on groups of related proteins, learning evolutionary constraints and relationships.
According to the developers, OpenProtein.AI is an open-ended toolbox, as the platform isn’t tied to a single protein function or class. Instead, the models are trained to understand proteins across the “whole space of possible proteins.” Researchers can adapt the workflows to their own needs, whether working on enzymes, antibodies, or novel proteins.
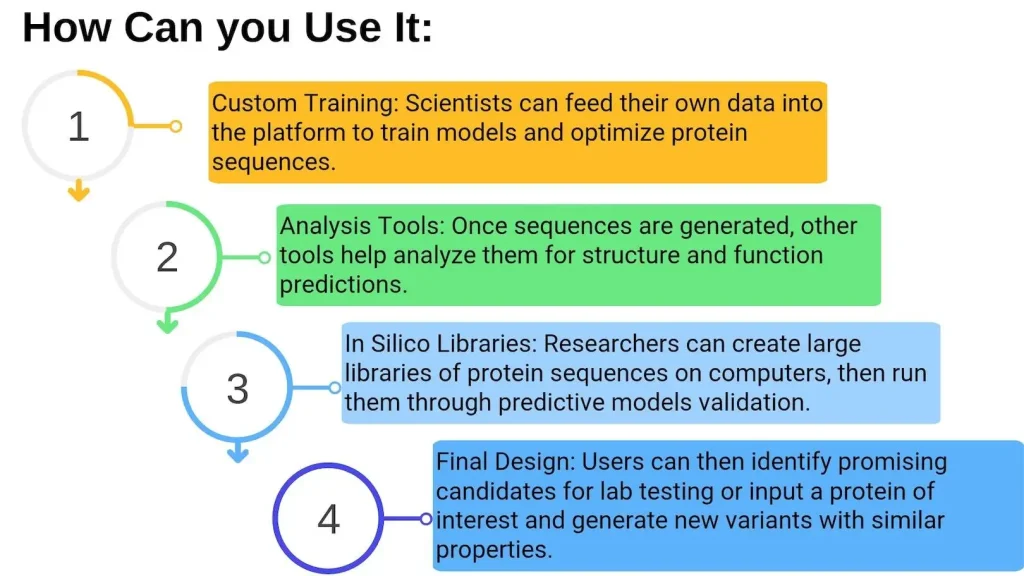
OpenProtein.AI’s user interface caters to all kinds of users. There is no need for any kind of programming or machine learning expertise; researchers can just upload their datasets using the web interface. This no-code approach makes the application more accessible to first-time users by enabling them to concentrate more on biology and less on the implementation aspect. However, the use of APIs is also an option for those who prefer coding.
Future-Oriented Vision of OpenProtein.AI: Updates and Collaborations
Bepler envisions a model that can describe not just the amino acid sequence, but also the functional context, like the enzymatic reaction a protein carries out. Lu’s perspective is to design a model that can predict and design proteins capable of engaging multiple biological mechanisms simultaneously while also changing their functions after binding, depending upon the context. This is to design proteins that operate closer to how natural proteins operate in real systems.
A model like this could design next-generation therapies that adapt to complex disease environments. In fact, Boehringer Ingelheim began using OpenProtein.AI’s platform in early 2025. This partnership embeds OpenProtien’s models directly into Boehringer’s workflows for engineering therapeutic proteins for cancer treatments, autoimmune diseases, and inflammatory conditions, meaning AI isn’t just a research tool anymore but a part of core drug development.
Moreover, PoET-2, which was released last year, builds on the original evolutionary transformer, outperforming much larger models while using only a fraction of the computing resources and experimental data, making it efficient enough to be used by ANY lab.
Conclusion
The startup sees its role as giving scientists the best possible tools to develop new therapies faster. OpenProtein.AI’s platform is designed to overcome the limitations faced by traditional experimental toolsets by embedding AI into everyday workflows. OpenProtein.AI’s commitment to free academic access and efficient models reflects this philosophy.
Article Source: Reference Article
Disclaimer:
The research discussed in this article was conducted and published by Insilico Medicine. CBIRT has no involvement in the research itself. This article is intended solely to raise awareness about recent developments and does not claim authorship or endorsement of the research.
Follow Us!
Learn More:
Saniya is a graduating Chemistry student at Amity University Mumbai with a strong interest in computational chemistry, cheminformatics, and AI/ML applications in healthcare. She aspires to pursue a career as a researcher, computational chemist, or AI/ML engineer. Through her writing, she aims to make complex scientific concepts accessible to a broad audience and support informed decision-making in healthcare.