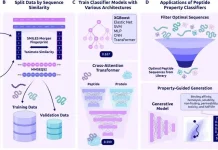
The researchers at the Mayo Clinic Cancer Center in Florida, in a recent study published in Nature Communications, validate the use of genetic sequencing to predict whether patients with gastric cancer would benefit from chemotherapy or immunotherapy.
Gastric cancer is one of the significant reasons for cancer-related deaths worldwide. Most patients with gastric cancer receive chemotherapy and, in certain cases, immunotherapy as part of their treatment regimen, according to Dr. Tae Hyun Dr. Hwang, chair at Mayo Clinic Cancer Center in Florida. These treatments, however, do not assist all patients. The researchers aimed to utilize genetic sequencing to create a model that might predict whether a patient would benefit from chemotherapy or immunotherapy.
Dr. Hwang and his colleagues created and executed a machine learning system that used genomic data from over 5,000 patients to create this model. The scientists then created a 32-gene molecular signature that might be used to guide patient treatment preferences.
Image Source: Development and validation of a prognostic and predictive 32-gene signature for gastric cancer.
Genomic profiling can guide clinical treatment, which can give prognostic and predictive information. In the case of gastric cancer, biomarkers that consistently predict patient response to chemotherapy and immune checkpoint suppression are absent.
The genetic landscape of gastric cancer has been clarified following recent large-scale next-generation sequencing and molecular profiling efforts. However, as functional assessments of gene-gene interaction and signaling system dynamics are not included in the mutational cataloging of cancer genomes, a potentially rich source of prognostic information is lost. The researchers previously created NTriPath, a machine learning technique that identifies predictive cancer-associated molecular pathways by combining pan-cancer somatic mutation data, gene-gene interaction networks, and pathway databases. The model NTriPath was employed to find predictive gene signatures for renal cell carcinoma, bladder carcinoma, head & neck squamous cell carcinoma, and melanoma.
In this investigation, the researchers employed their machine learning algorithm ‘NTriPath’ to find a 32-gene signature unique to stomach cancer. They discover four molecular subgroups that are predictive for survival using unsupervised clustering on expression levels of these 32 genes in malignancies from 567 individuals. The scientists then used a support vector machine with a linear kernel to develop a risk score that is prognostic for five-year overall survival, which they subsequently validated using three different datasets. The molecular subtypes also predict response to adjuvant 5-fluorouracil and platinum treatment following gastrectomy, as well as immune checkpoint inhibitors, in patients with metastatic or recurrent illness, according to the researchers.
To summarise, the researchers show that the 32-gene signature is a promising prognostic and predictive biomarker for guiding the clinical management of gastric cancer patients and that it should be confirmed prospectively using large patient cohorts.
The researchers discovered that the 32-gene signature provided prognostic information and predicted whether the patient benefits from chemotherapy and immunotherapy. To the researcher’s amazement, the 32-gene signature found may predict a patient’s immunotherapy response. Establishing meaningful biomarkers for immunotherapy response in individuals with gastric cancer has proven difficult.
The 32-gene molecular signature, according to Dr. Hwang, still has to be prospectively validated, but he hopes it may eventually be able to identify individuals who will show response to chemotherapy and immunotherapy. Similarly, the authors would also be able to identify individuals who are unlikely to benefit from chemotherapy or immunotherapy, avoiding the potentially harmful side effects of these treatments.
To make biomarkers more accessible and easy to deploy in the clinical context, the team is developing novel assays based on the expression level of single or multiple genes.
“We are working on artificial intelligence algorithms that utilize diagnostic histopathology images to identify patients most likely to derive benefit from immunotherapy,” expresses Dr. Hwang. “We are also studying the molecular mechanisms of immunotherapy resistance made available by the machine learning and artificial intelligence approaches that we have developed in our lab.”
Story Source: Cheong, JH., Wang, S.C., Park, S. et al. Development and validation of a prognostic and predictive 32-gene signature for gastric cancer. Nat Commun 13, 774 (2022). https://doi.org/10.1038/s41467-022-28437-y
https://newsnetwork.mayoclinic.org/discussion/mayo-clinic-researchers-develop-model-to-predict-treatment-response-in-gastric-cancer/
Dr. Tamanna Anwar is a Scientist and Co-founder of the Centre of Bioinformatics Research and Technology (CBIRT). She is a passionate bioinformatics scientist and a visionary entrepreneur. Dr. Tamanna has worked as a Young Scientist at Jawaharlal Nehru University, New Delhi. She has also worked as a Postdoctoral Fellow at the University of Saskatchewan, Canada. She has several scientific research publications in high-impact research journals. Her latest endeavor is the development of a platform that acts as a one-stop solution for all bioinformatics related information as well as developing a bioinformatics news portal to report cutting-edge bioinformatics breakthroughs.