Researchers have discovered molecular fingerprints that lead to biological causes of disease by analyzing individual cardiac cells from heart failure patients.
Heart failure is an increasingly important public health issue because it comprises a diverse range of clinical characteristics that are all related to reduced heart contractile function.
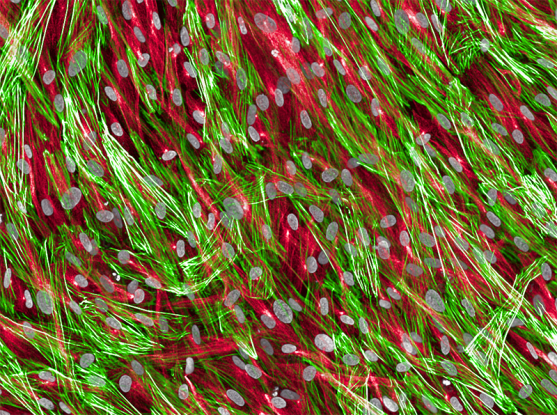
Fibroblasts, a kind of connective tissue cell in the heart, showing cell nuclei (gray), tubulin (red), and actin (green).
Image Source: https://www.broadinstitute.org/news/single-cell-map-heart-failure-suggests-possible-therapeutic-targets
By performing single-nucleus RNA sequencing on nearly 600,000 nuclei in left ventricle samples from 16 non-failing hearts and 11 hearts with dilated cardiomyopathy and 15 hearts with hypertrophic cardiomyopathy, this study identifies extensive molecular alterations in failing hearts at single-cell resolution.
The dilated and hypertrophic cardiomyopathies (DCM and HCM), which both impair the heart’s ability to pump blood, are two significant causes of heart failure. Researchers from the Precision Cardiology Lab (PCL) of the Broad Institute of MIT, Harvard, and Bayer have created detailed maps of a variety of cell types in the heart that are involved in these two conditions.
The team’s results, which were released in the journal Nature, point to particular cell types and biological processes that novel treatments might target. Heart failure is one of the most common reasons for hospitalization in the US. Existing medications are limited, and many patients eventually pass away from heart failure.
The researchers employed single-nuclei RNA sequencing (RNA-seq), which reveals which genes are activated and at what amounts in individual cells, to produce their cellular maps. They examined cardiac tissue from individuals with advanced heart failure and discovered that DCM and HCM patients had comparable gene expression profiles even though their heart cells expressed distinct genes from non-failing hearts. Additionally, certain cardiomyopathy patients showed a distinct collection of fibroblasts, or connective tissue cells, which the researchers believe may be involved in tissue scarring in heart failure and may be a potential target for treatment in the future.
This research is the product of close cooperation with the team led by Ken Margulies, a professor of medicine and heart failure specialist at the University of Pennsylvania.
This study builds on a prior attempt to catalog individual cells in the healthy human heart. Researchers may be able to uncover markers that would enable them to distinguish between different disease types and predict clinical outcomes by mapping the cells involved in various types of heart failure.
According to Patrick Ellinor, who led the team and is a member of the Broad Institute, director of the Demoulas Center for Cardiac Arrhythmias at Massachusetts General Hospital, and professor at Harvard Medical School, almost all types of heart failure are currently treated the same way, regardless of their underlying causes. The scientists’ main query was whether there were any cell populations or gene expression differences between healthy and diseased individuals in those who had a condition. And certainly, the researchers discovered that some diseased patients had genes indicating highly active cardiac fibroblasts.
The data for this investigation were generated, analyzed, and validated by scientists from Bayer and Broad. Collaboration between academics and the pharmaceutical business at this level is highly uncommon, said Carla Klattenhoff, senior director and head of the joint Precision Cardiology Laboratory of Bayer and the Broad. This map is a fantastic resource for the field of cardiology.
Ulrich Nielsch, head of Bayer’s Therapeutic Area 1, stated that this is a significant scientific contribution to increasing our understanding of disease and enabling the delivery of precision medicines for cardiology.
Analyzing individual cardiac cells
In contrast to healthy hearts, failing hearts have distinct gene expression profiles or transcriptomes; this research only produced a single genetic fingerprint for the entire heart. Contrarily, Ellinor’s team used computational techniques to differentiate transcriptional fingerprints by cell type using single nucleus RNA-seq. Additionally, they were able to detect signs of uncommon cell types whose signals could have been masked by bulk analysis.
The researchers examined DCM and HCM, two conditions that cause heart failure in various ways. The left ventricle of the heart enlarges, and its walls thin in DCM, whereas they stiffen and thicken in HCM. Surprisingly, the scientists discovered that despite these variations, the situations share the same transcriptional fingerprint. This discovery may enable doctors to more accurately differentiate between kinds and stages of heart failure and determine the best course of treatment.
Additionally, Ellinor’s team discovered that people with cardiomyopathy and healthy individuals have different ratios of specific heart cell types. Compared to non-failing hearts, failing hearts exhibited fewer muscle cells but more fibroblasts, which may be a sign of scar tissue. The researchers also identified a distinct population of these fibroblasts that was restricted to failing hearts. As the study’s first author and a computational scientist at the Broad Institute, Mark Chaffin, stated, it was remarkable how particular the transcriptional profile of these cells actually was. In contrast to almost none in hearts that were not failing, there were thousands of these nuclei in cardiomyopathy hearts.
The scientists investigated the role of important genes that set this group of fibroblasts apart from typical cardiac fibroblasts using CRISPR screens. In order for cardiac fibroblasts to change from a dormant to an active state, in which cells generate scar tissue that can impair heart pumping, it was discovered that multiple genes were required. According to Chaffin, these genes may serve as potential targets for fibrosis treatments in the future.
By examining biomarkers of particular types of fibroblasts or higher levels of scarring in the heart, the researchers aim to one day discover a method to identify heart failure in its earliest stages in patients. The group has begun examining whether they can find biomarkers of active fibroblasts in blood in order to meet this objective.
Additionally, Ellinor’s group looks forward to creating single-cell maps of several cardiovascular diseases, such as ischemic heart disease and atrial fibrillation.
Story Source: Chaffin, M., Papangeli, I., Simonson, B. et al. Single-nucleus profiling of human dilated and hypertrophic cardiomyopathy. Nature (2022). https://doi.org/10.1038/s41586-022-04817-8
https://www.broadinstitute.org/news/single-cell-map-heart-failure-suggests-possible-therapeutic-targets
Learn More About Bioinformatics:
Top Bioinformatics Books ↗
Learn more to get deeper insights into the field of bioinformatics.
Top Free Online Bioinformatics Courses ↗
Freely available courses to learn each and every aspect of bioinformatics.
Latest Bioinformatics Breakthroughs ↗
Stay updated with the latest discoveries in the field of bioinformatics.
Dr. Tamanna Anwar is a Scientist and Co-founder of the Centre of Bioinformatics Research and Technology (CBIRT). She is a passionate bioinformatics scientist and a visionary entrepreneur. Dr. Tamanna has worked as a Young Scientist at Jawaharlal Nehru University, New Delhi. She has also worked as a Postdoctoral Fellow at the University of Saskatchewan, Canada. She has several scientific research publications in high-impact research journals. Her latest endeavor is the development of a platform that acts as a one-stop solution for all bioinformatics related information as well as developing a bioinformatics news portal to report cutting-edge bioinformatics breakthroughs.