Yale scientists report in the journal Nature Biotechnology this week that a novel data analysis technique they created has uncovered the particular immune cell types linked to an elevated risk of mortality from COVID-19.
A variety of technologies provide high-throughput biological data by measuring dozens to tens of thousands of features in millions of cells collected from big patient cohorts. More powerful computer algorithms are required to derive biological insights. The researchers introduce Multiscale PHATE, a technique for learning abstracted biological traits that are directly predictive of illness outcome that sweeps overall levels of data granularity.
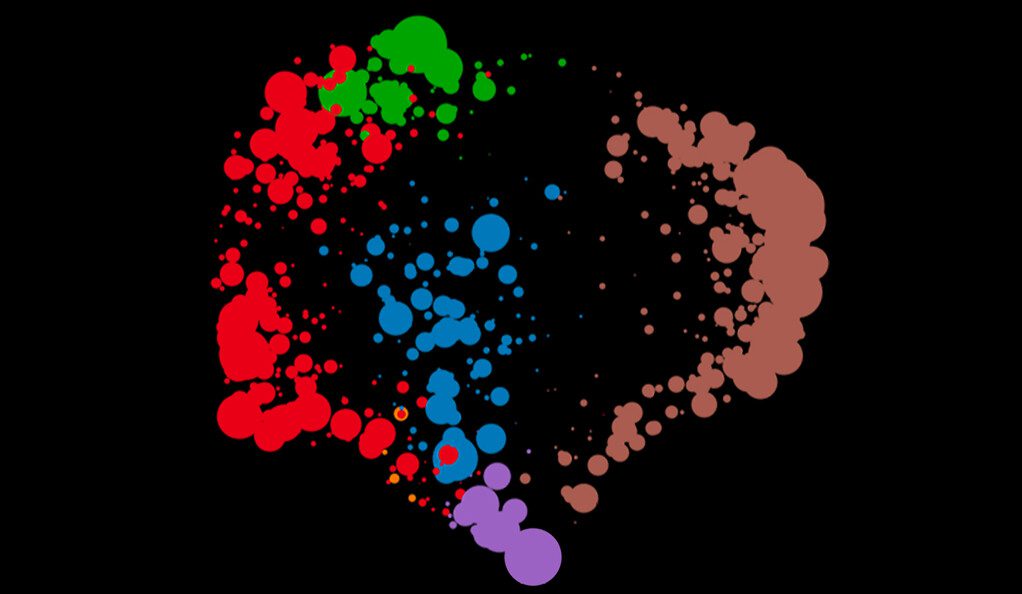
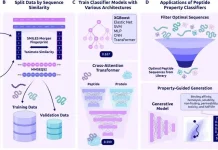
Image Source: Multiscale PHATE identifies multimodal signatures of COVID-19.
T cells and antibody-producing B cells in the immune system have been shown to give widespread protection against infections like SARS-CoV-2, the virus that causes COVID-19. In addition, large-scale data analysis involving millions of cells has provided scientists with a wide picture of the immune system’s reaction to this virus. They have discovered, however, that some immune cell responses, even those from normally protective cell types, can occasionally cause fatal inflammation and death in patients.
Other data processing technologies that enable research down to the level of single cells have provided scientists with some information concerning the causes of severe COVID instances. However, such targeted views frequently lack the context of specific cell groupings that may result in better or worse results.
Researchers can pass through all resolutions of data, from millions of cells to a single cell, in minutes using the Multiscale PHATE tool, a machine learning technology created at Yale. The technique is based on the PHATE algorithm, which was developed by the group of Smita Krishnaswamy, associate professor of genetics and computer science. This technique also addresses many of the flaws of existing data visualization tools.
“Machine learning algorithms typically focus on a single resolution view of the data, ignoring information that can be found in other more focused views,” said Manik Kuchroo, a doctoral candidate at Yale School of Medicine who helped develop the technology and is co-lead author of the paper. “For this reason, we created Multiscale PHATE which allows users to zoom in and focus on specific subsets of their data to perform more detailed analysis.”
Kuchroo, who is Krishnaswamy’s lab member, utilized the new technology to examine 55 million blood cells from 163 COVID-19 patients hospitalized to Yale New Haven Hospital. They discovered that high amounts of T cells appear to protect against bad outcomes, but high levels of two white blood cell types, granulocytes, and monocytes were linked to greater rates of death.
When the researchers delved down to a more granular level and clustered TH17, a helper T cell, with the immune system cells IL-17 and IFNG they observed that TH17 was associated with higher mortality.
According to the researchers, they were able to predict whether a patient would live or die with 83 percent accuracy by analyzing the number of these cells in the blood.
“We were able to rank order risk factors of mortality to show which are the most dangerous,” Krishnaswamy said.
Multiscale PHATE is greatly customizable as it can be used on various data types, such as flow cytometry, scRNA-seq, scATAC-seq, and clinical variables. According to the authors, the new data analysis method may theoretically be used to fine-tune risk assessment in a variety of conditions.
Story Source: Kuchroo, M., Huang, J., Wong, P. et al. Multiscale PHATE identifies multimodal signatures of COVID-19. Nat Biotechnol (2022). https://doi.org/10.1038/s41587-021-01186-x
Background Image Description: Multiscale PHATE plot of immune cells of COVID-19 patients shows a coarse-grained visualization and clustering of the cells allowing for detection of predictive, prognostic populations at fine granularity.
Background Image Source: https://news.yale.edu/2022/02/28/yales-new-data-analysis-tool-uncovers-important-covid-19-clues
Dr. Tamanna Anwar is a Scientist and Co-founder of the Centre of Bioinformatics Research and Technology (CBIRT). She is a passionate bioinformatics scientist and a visionary entrepreneur. Dr. Tamanna has worked as a Young Scientist at Jawaharlal Nehru University, New Delhi. She has also worked as a Postdoctoral Fellow at the University of Saskatchewan, Canada. She has several scientific research publications in high-impact research journals. Her latest endeavor is the development of a platform that acts as a one-stop solution for all bioinformatics related information as well as developing a bioinformatics news portal to report cutting-edge bioinformatics breakthroughs.
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar