Large-scale proteome and genomic investigations play a crucial role in understanding the molecular causes of cancer. The National Cancer Institute’s Clinical Proteomic Tumor Analysis Consortium (CPTAC) has developed an invaluable resource of data, cProSite (Cancer Proteogenomic Data Analysis Site), a user-friendly website designed for biologists and clinicians, offering online analysis of genomics, proteomics, and phosphoproteomics using CPTAC data. cProSite enables the comparison and correlation of proteins, mRNAs, and phosphorylation sites in tumors and normal tissues, providing insights into potential drug targets at various molecular levels.
cProSite: A Powerful Analytical Tool
cProSite is an interactive database that allows researchers to explore the expression and correlation of proteins, proteomic phosphorylations, and mRNA levels between tumor types and normal tissues. Unlike traditional analytical methods, cProSite offers several advantages, including faster online analysis, a user-friendly environment, and reduced reliance on bioinformatics expertise.
cProSite integrates proteomics and phosphoproteomics data from the Proteomic Data Commons (PDC) and CPTAC portal, which have undergone uniform processing through the CPTAC Common Data Analysis Pipeline (CDAP). The website also incorporates CPTAC mRNA data from the Genomic Data Commons Data Portal. These datasets ensure comparability across different samples and cancer types. The web platform utilizes an SQLite table database and employs React, a JavaScript library, for the front-end interface. NodeJS is used for efficient querying of the database on the platform’s backend.
Analyzing Protein Abundance, Phosphorylation, and mRNA Levels
cProSite provides comprehensive analysis capabilities for protein abundance and phosphorylation levels in tumors and normal tissues. It enables users to explore protein abundance differences within specific tumor types as well as across different tumor types. For example, users can examine the log2 fold changes in protein abundance of specific proteins, such as caveolin 1 (CAV1), across various tumor types. The results unveil intriguing insights into the downregulation of CAV1 in most tumor types except for kidney cancer.
Moreover, cProSite allows the analysis of phosphorylation levels at individual phosphorylation sites within different tumor types. The platform presents a heatmap-like visualization of phosphorylation site levels, enabling users to observe the log2 fold changes in phosphorylation sites for specific proteins like CAV1. Additionally, by introducing the concept of phosphorylation level per protein abundance level, researchers can assess the actual phosphorylation level of a protein relative to its abundance.
The website also facilitates the comparison of mRNA levels between tumors and normal tissues. Users can explore the expression of the same gene at both the protein and RNA levels, gaining a comprehensive view of gene expression in cancer.
Investigating Correlations between Protein Abundance and RNA Expression
Understanding the relationship between protein abundance and mRNA expression is essential for unraveling gene expression control mechanisms. The cProSite platform offers sophisticated tools to evaluate correlations between protein abundance, including inter-protein, protein-phosphorylation site, and phosphorylation site-phosphorylation site associations. Researchers can explore the correlation between protein levels and the corresponding mRNA expression, shedding light on the dynamic transitions and regulatory mechanisms involved.
For instance, the analysis reveals that some crucial proteins involved in the G2/M phase transition of the cell cycle exhibit low mRNA levels in normal tissues. However, both mRNA and protein levels are significantly enhanced in tumors, indicating a strong coupling between mRNA transcription and active protein synthesis in cancer cells.
Clinical Data Validation and Experimental Confirmation
cProSite serves as a valuable resource for clinical data validation. Researchers can compare protein abundance changes between tumors and normal tissues, providing indicators for clinical validation. By utilizing cProSite, researchers have successfully validated the expression of specific proteins, such as the receptor for hyaluronic acid-mediated motility (RHAMM), which is linked with poor prognosis and metastasis in non-small cell lung carcinoma (NSCLC). The platform’s protein abundance comparison confirmed the high expression of RHAMM in tumors, consistent with tissue microarray analysis.
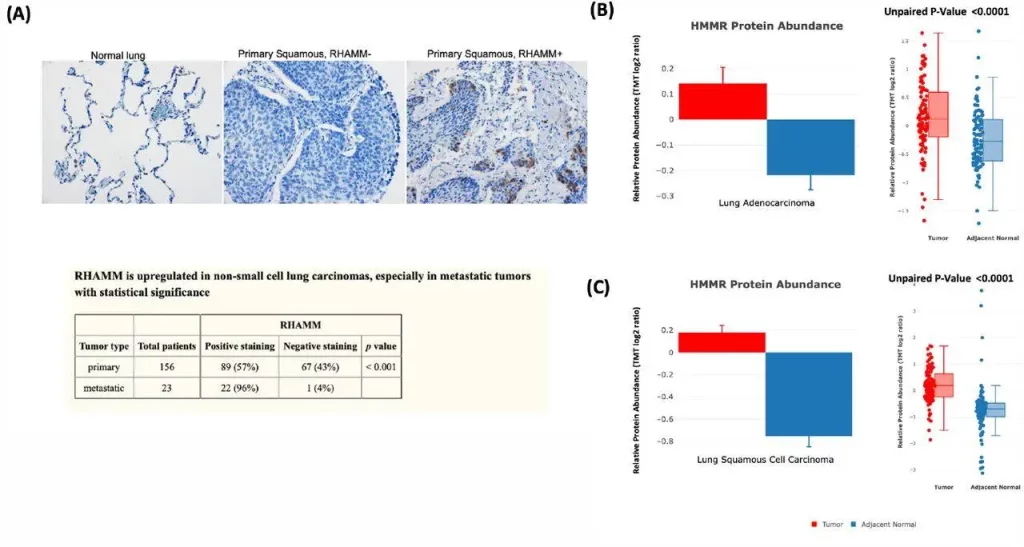
Image Source: https://doi.org/10.1101/2023.06.10.543932
Additionally, cProSite has been employed to investigate the phosphorylation sites of proteins involved in cell cycle progression. By examining the correlation between CDK1 kinase activity and phosphorylation levels, researchers have identified potential prognostic markers. Experimental validation of these phosphorylation sites further confirmed their functional role in cell cycle regulation.
Conclusion
cProSite represents a significant advancement in the field of genomics and proteomics analysis. It provides a user-friendly and efficient web-based platform for researchers, biologists, and clinicians to explore molecular alterations in tumors and normal tissues. Its features include protein abundance analysis, phosphorylation site comparison, and correlation analysis between proteins and genes.
Article Source: Reference Paper | cProSite: Website
Important Note: bioRxiv releases preprints that have not yet undergone peer review. As a result, it is important to note that these papers should not be considered conclusive evidence, nor should they be used to direct clinical practice or influence health-related behavior. It is also important to understand that the information presented in these papers is not yet considered established or confirmed.
Learn More:
Dr. Tamanna Anwar is a Scientist and Co-founder of the Centre of Bioinformatics Research and Technology (CBIRT). She is a passionate bioinformatics scientist and a visionary entrepreneur. Dr. Tamanna has worked as a Young Scientist at Jawaharlal Nehru University, New Delhi. She has also worked as a Postdoctoral Fellow at the University of Saskatchewan, Canada. She has several scientific research publications in high-impact research journals. Her latest endeavor is the development of a platform that acts as a one-stop solution for all bioinformatics related information as well as developing a bioinformatics news portal to report cutting-edge bioinformatics breakthroughs.