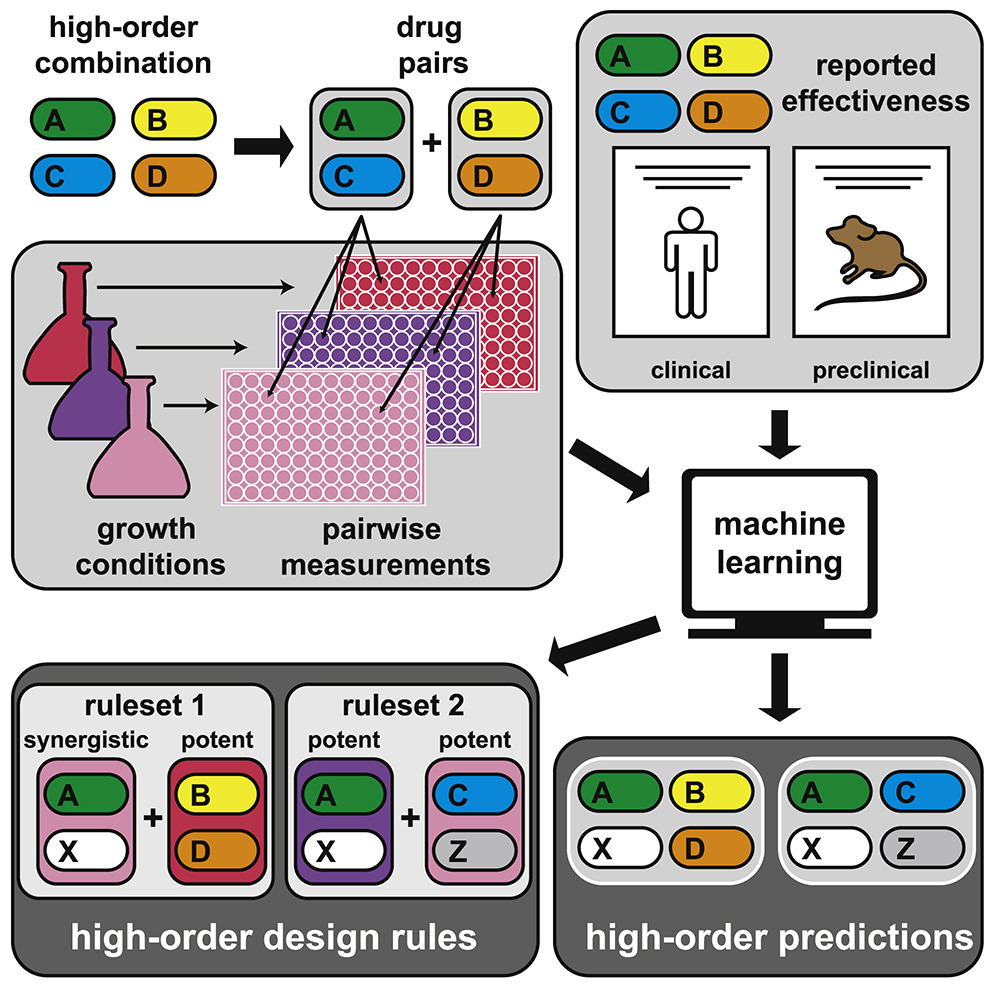
A challenge to tuberculosis treatment regimen design is the need to combine three or more antibiotics. The study shows how machine learning can provide a data-driven perspective on this challenge, so researchers can consider new drug combinations to provide better treatments. The researchers believe this new approach will increase scientists’ ability to determine which drug combinations will be most successful in combating tuberculosis, the world’s second leading infectious killer.
Think of 20 new compounds that have shown some efficacy against a disease like tuberculosis (TB) which affects 10 million people globally and kills 1.5 million every year. TB bacteria behave differently in different environments—and can even evolve to become drug-resistant—so patients will need a combination of three or four drugs to fight the disease effectively.
What factors should be considered when deciding which drugs to test together? The combination of 20 compounds in three- and four-drug combinations offers nearly 6,000 combinations. A recent study by Tufts University researchers, published in Cell Reports Medicine, demonstrated how machine learning could provide a data-driven perspective on this challenge so that researchers can consider new drug combinations at a greater level without testing each combination on animal models.
According to Bree Aldridge, associate professor of molecular biology and microbiology at Tufts University School of Medicine and of biomedical engineering at the School of Engineering and a member of the immunology and molecular microbiology program faculty at the Graduate School of Biomedical Sciences, the framework can be used to predict how effective treatments will be when we move from lab testing to mouse tests, which is an important step before moving on to human trials.
This method makes it easier to determine which multi-drug treatments should be moved into preclinical trials. Moreover, it provides interpretable rules that researchers and drug developers can use to design optimal combinations of drugs.
According to Jonah Larkins-Ford, GBS22, co-first author on the paper and a principal scientist at MarvelBiome, now instead of guessing and testing every possible combination in animals, the research provides empirical evidence about how drug combinations should be designed. With these concrete rules, one can identify new three- or four-drug cocktails to improve the treatment of tuberculosis with deliberate, data-driven design.
Aldridge’s lab developed and uses DiaMOND, or diagonal measurement of n-way drug interactions, which is a method developed and used to systemically study pairwise and high-order drug combination interactions in order to identify shorter, more efficient treatment regimens for tuberculosis and other diseases that require combination drug therapy, such as cancer. The researchers believe that by using the design rules established in the new study, their new system can speed up the process by which scientists determine which drug combinations will most effectively treat tuberculosis.
Learn More:
Top Bioinformatics Books ↗
Learn more to get deeper insights into the field of bioinformatics.
Top Free Online Bioinformatics Courses ↗
Freely available courses to learn each and every aspect of bioinformatics.
Latest Bioinformatics Breakthroughs ↗
Stay updated with the latest discoveries in the field of bioinformatics.
Dr. Tamanna Anwar is a Scientist and Co-founder of the Centre of Bioinformatics Research and Technology (CBIRT). She is a passionate bioinformatics scientist and a visionary entrepreneur. Dr. Tamanna has worked as a Young Scientist at Jawaharlal Nehru University, New Delhi. She has also worked as a Postdoctoral Fellow at the University of Saskatchewan, Canada. She has several scientific research publications in high-impact research journals. Her latest endeavor is the development of a platform that acts as a one-stop solution for all bioinformatics related information as well as developing a bioinformatics news portal to report cutting-edge bioinformatics breakthroughs.
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar