New research from University of Oxford revealed that superbug Pseudomonas develops resistance to a common 'last-resort' antibiotic much faster than expected.
Recent research published in Cell Reports this week shows how populations of the bacterium Pseudomonas react to treatment with Colistin, a ‘last resort’ antibiotic for patients with multidrug-resistant infections.
Antibiotics are essential for human health because they help fight infection, but bacteria can develop resistance to the antibiotics that patients rely on. Antibiotic-resistant diseases now kill over a million people per year around the world.
Colistin has emerged as a critical last line of defense for treating infections caused by antibiotic-resistant gram-negative bacteria such as Escherichia coli, Klebsiella pneumonia, Pseudomonas aeruginosa, and Acinetobacter baumanii, however, colistin resistance is poorly understood.
With a limited supply of ‘last-resort’ antibiotics, University of Oxford researchers are looking into the processes that cause the rise and fall of resistance in common bacterial pathogen populations, which is crucial in combating antimicrobial resistance (AMR).
According to Professor Craig MacLean, Department of Biology, University of Oxford, the research has discovered that a gene implicated in last-resort antibiotic resistance mutates at an extremely high rate, allowing bacteria to rapidly evolve antibiotic resistance.
Further, Professor Craig MacLean explained that the findings show that selective pressures induced by this gene’s interaction with the immune system may have aided the evolution of an extra-fast mutation rate, which is rapidly developing to make bacteria antibiotic-resistant.
Pseudomonas is a bacteria that causes pneumonia in hospital patients. Over 900 populations of Pseudomonas were cultivated and treated with Colistin by the researchers. The researchers were able to determine how rapidly distinct populations gained resistance to the antibiotic and the genetic mutation that caused resistance by counting bacteria and sequencing their genomes.
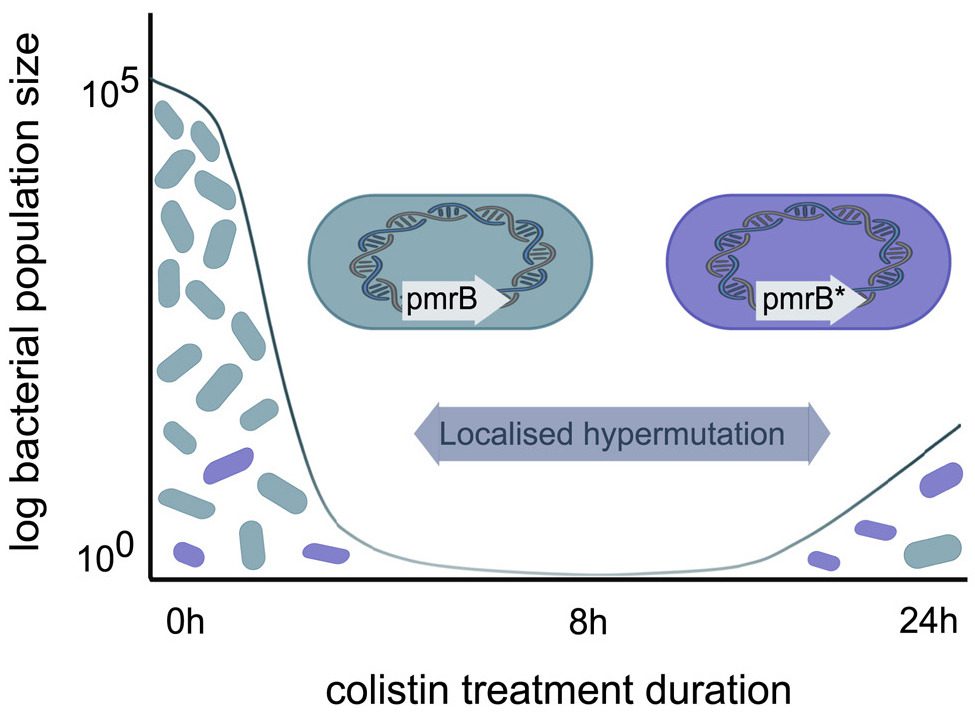
The findings revealed that Pseudomonas infections developed resistance to the last-resort antibiotic swiftly because of the gene that mutates 1,000 times faster than the ‘normal’ background rate of mutation. The bacteria were able to evolve their resistance to Colistin due to mutations in this gene, known as pmrB.
According to researchers, the pmrB gene is linked to the human immune system, which could explain why mutations happen so quickly. A high mutation rate would aid the bacterium’s ability to adapt to variable immune system changes.
Despite the fact that the bacteria developed resistance to Colistin at a considerably faster rate than planned, the study had a favorable effect. Because of the rapid mutation rate, pathogen populations quickly lost resistance when the antibiotic was removed.
The researchers now intend to expand their research to see if additional Pseudomonas characteristics are involved in enabling such high levels of antibiotic resistance.
Story Source: Kapel, N., Caballero, J. D., & MacLean, R. C. (2022). Localized <em>pmrB</em> hypermutation drives the evolution of colistin heteroresistance. Cell Reports, 39(10). https://doi.org/10.1016/j.celrep.2022.110929
https://www.ox.ac.uk/news/2022-06-07-common-drug-resistant-superbug-develops-fast-resistance-last-resort-antibiotic
Dr. Tamanna Anwar is a Scientist and Co-founder of the Centre of Bioinformatics Research and Technology (CBIRT). She is a passionate bioinformatics scientist and a visionary entrepreneur. Dr. Tamanna has worked as a Young Scientist at Jawaharlal Nehru University, New Delhi. She has also worked as a Postdoctoral Fellow at the University of Saskatchewan, Canada. She has several scientific research publications in high-impact research journals. Her latest endeavor is the development of a platform that acts as a one-stop solution for all bioinformatics related information as well as developing a bioinformatics news portal to report cutting-edge bioinformatics breakthroughs.
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar











