Novel discovery by scientists from UT Southwestern University identifies co-component signal transduction systems in gut bacteria that could help fight gastrointestinal infections.
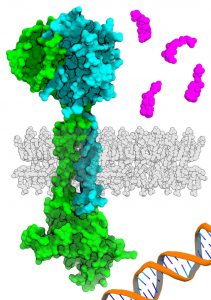
Image Source: https://www.utsouthwestern.edu/newsroom/articles/year-2022/june-gut-bacteria.html
Scientists used artificial intelligence to identify a new family of enteric bacteria sensing genes that are related by structure and likely function but not genetic sequence. The research, which was published in PNAS, provides a new method for determining the function of genes in unrelated species and may help develop new treatments for intestinal bacterial infections.
According to Kim Orth, Ph.D., Professor of Molecular Biology and Biochemistry, who co-led the study with Lisa Kinch, Ph.D., a bioinformatics specialist in the Department of Molecular Biology, the researchers found similarities in these proteins as opposed to how it’s normally done. Rather than using sequence, Lisa looked for matches in their structure.
Dr. Orth’s work has long focused on determining how bacteria in estuaries and the oceans cause diseases. In 2016, Dr. Orth and her coworkers employed biophysics to define the structure of the VtrA and VtrC complex, two proteins that cooperate in the Vibrio parahaemolyticus bacterial species. The researcher then discovered that VtrA/VtrC complex in V. parahaemolyticus, which frequently causes food poisoning from contaminated shellfish – senses bile from the bacterial cell surface, sending a signal to start a chemical cascade that causes this microbe to invade the intestinal cells of its human host.
It was unknown whether a homolog for VtrC existed in this or any other bacteria, despite the fact that ToxR, a protein from a related bacteria called Vibrio cholerae that causes cholera, shared certain structural similarities with VtrA.
We had never seen anything like VtrC. But, we thought, other proteins like it must exist.
Dr. Kinch
Without being able to find any known genes with sequence identities similar to VtrC, the researchers utilized a program called AlphaFold, which was only launched two years ago. On the basis of genetic sequence that codes for specific proteins, this artificial intelligence program can predict the structure of such proteins with high accuracy—information that was previously only obtained through time-consuming laboratory effort.

Image Source: Co-component signal transduction systems: Fast-evolving virulence regulation cassettes discovered in enteric bacteria
Despite the lack of any identifiable genetic sequences between the two proteins, AlphaFold demonstrated that a protein termed ToxS in V. cholerae shares a great deal of structural similarity with VtrC. The researchers discovered homologs of VtrC in several other enteric bacterial species that cause human disease when they looked for proteins in other organisms with comparable structural characteristics. These bacteria include Yersinia pestis, which causes the bubonic plague, and Burkholderia pseudomallei (which causes a tropical infection called melioidosis). These VtrC homologs all appear to cooperate with proteins that share structural similarities with VtrA, indicating that they may play the same functions as in V. parahaemolyticus.
According to Dr. Orth, these structural similarities could potentially result in pharmaceuticals that address diseases brought on by various infectious organisms that employ comparable pathogenic methods.
Dr. Orth is a W.W. Caruth, Jr. Scholar in Biomedical Research and a Howard Hughes Medical Institute Investigator who holds the Earl A. Forsyth Chair in Biomedical Science. This is the first publication of the senior author in the journal PNAS, and she has been a National Academy of Sciences member since 2020.
Qian Cong, Ph.D., a Southwestern Medical Foundation Scholar in Biomedical Research, and Jananee Jaishankar, Ph.D., are two more UTSW researchers who made contributions to this study.
Conclusion of the study
The study reveals that the heterodimeric VtrA/VtrC periplasmic bile acid receptor governing the Vibrio parahaemolyticus type 3 secretion system 2 is a distant homolog of the ToxR/ToxS master regulator of virulence and has evolved past confident sequence recognition. The researchers observed a VtrC-like lipocalin fold for the ToxS periplasmic domain as well as other discovered periplasmic sensor components by making use of the most recent machine learning techniques for structure prediction (AlphaFold). The different evolution of VtrA/VtrC-like co-component signal transduction systems is supported by this structural prediction, which also raises the possibility that lipid sensing may play a role in controlling virulence in enteric bacteria.
Story Source: N., K. L., Qian, C., Jananee, J., & Kim, O. (2022). Co-component signal transduction systems: Fast-evolving virulence regulation cassettes discovered in enteric bacteria. Proceedings of the National Academy of Sciences, 119(24), e2203176119. https://doi.org/10.1073/pnas.2203176119 https://www.utsouthwestern.edu/newsroom/articles/year-2022/june-gut-bacteria.html
Learn More About Bioinformatics:
Top Bioinformatics Books ↗
Learn more to get deeper insights into the field of bioinformatics.
Top Free Online Bioinformatics Courses ↗
Freely available courses to learn each and every aspect of bioinformatics.
Latest Bioinformatics Breakthroughs ↗
Stay updated with the latest discoveries in the field of bioinformatics.
Dr. Tamanna Anwar is a Scientist and Co-founder of the Centre of Bioinformatics Research and Technology (CBIRT). She is a passionate bioinformatics scientist and a visionary entrepreneur. Dr. Tamanna has worked as a Young Scientist at Jawaharlal Nehru University, New Delhi. She has also worked as a Postdoctoral Fellow at the University of Saskatchewan, Canada. She has several scientific research publications in high-impact research journals. Her latest endeavor is the development of a platform that acts as a one-stop solution for all bioinformatics related information as well as developing a bioinformatics news portal to report cutting-edge bioinformatics breakthroughs.












