According to recent research published in Nature Communications, a new AI algorithm creates novel compounds based on desired properties.
Researchers are searching for innovative chemicals in a variety of fields, including health, battery research, and materials science. They can typically forecast the necessary chemical and physical features in great precision during this process, down to the atomic level. However, the overall number of possible chemical compounds is so large that finding the suitable material would take years. An interdisciplinary research group at Technische Universität Berlin’s Berlin Institute for the Foundations of Learning and Data (BIFOLD) has developed an artificial intelligence algorithm to implement inverse chemical design and generate targeted molecules based on their desired properties.
The quest for appropriate compounds for specific medicinal or industrial purposes is a time-consuming and costly endeavor. “Theoretically, there are an infinite number of structures that may exist. Only a small percentage of them, however, have the chemical or physical qualities necessary for a given application,” says Dr. Kristof Schütt, a BIFOLD Junior Fellow at TU Berlin.
In recent years, many AI-based approaches for predicting the chemical characteristics and energy states of specified molecules have been created. However, even with these effective approaches, finding molecules with specified features has proven challenging in reality, as it requires sifting through a large number of possibilities.
Reversing the link between structure and property
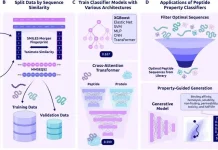
As a result, the BIFOLD research group focuses on inverse molecular design, in which the structure-property link is inverted. The properties define the structure rather than the structure determining the properties. The task entails creating molecular structures from scratch that correspond to a set of attributes. The artificial intelligence method is built on a deep generative neural network that incorporates previous knowledge of primary physical conditions. To understand the complicated links between chemical structures and their characteristics, the network only takes a few thousand example molecules.
“The user can then specify various property values, and the generative neural network suggests a manageable number of suitable molecules and compounds. Only these candidates have to be investigated by the chemists,” expresses Schütt.
According to the researchers, inverse chemical design works even when the intended property values are only partially covered by the known sample of molecules.
In many functional domains, the multidisciplinary research team believes that using such algorithms in conjunction with other AI-driven methodologies and quantum chemistry methods can dramatically expedite the hunt for novel chemicals and materials.
“I see enormous potential here if both the design of the molecules and their analysis and simulation are supported by artificial intelligence methods. This could help in the development of drugs, for example, or accelerate the search for novel materials for batteries and solar cells,” said Klaus-Robert Müller, BIFOLD co-director and professor of machine learning at TU Berlin.
Story Source: Gebauer, N.W.A., Gastegger, M., Hessmann, S.S.P. et al. Inverse design of 3d molecular structures with conditional generative neural networks. Nat Commun 13, 973 (2022). https://doi.org/10.1038/s41467-022-28526-y
https://www.tu.berlin/en/about/profile/press-releases-news/2022/februar/function-determines-form/
Dr. Tamanna Anwar is a Scientist and Co-founder of the Centre of Bioinformatics Research and Technology (CBIRT). She is a passionate bioinformatics scientist and a visionary entrepreneur. Dr. Tamanna has worked as a Young Scientist at Jawaharlal Nehru University, New Delhi. She has also worked as a Postdoctoral Fellow at the University of Saskatchewan, Canada. She has several scientific research publications in high-impact research journals. Her latest endeavor is the development of a platform that acts as a one-stop solution for all bioinformatics related information as well as developing a bioinformatics news portal to report cutting-edge bioinformatics breakthroughs.
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar
- Dr. Tamanna Anwar