There is a question that has quietly haunted drug discovery for decades, one that sounds almost too basic to be a problem. A molecule finds its target protein. It docks in precisely. But then what? Does anything actually happen inside the cell? For a surprisingly long time, computational tools could answer the first part of that question reasonably well, while the second part remained largely out of reach. A research team at the Korea Advanced Institute of Science and Technology (KAIST), South Korea, took that second question seriously and developed GPCRact, an AI framework published in January 2026 in Briefings in Bioinformatics that reframes how to predict drug activity from the ground up.
The Problem With Just Knowing a Drug Binds
In the cell, G-protein-coupled receptors, or GPCRs, respond to a variety of external stimuli, including hormones, neurotransmitters, and drugs. After stimulus recognition, GPCRs communicate the stimuli internally and initiate a biological response. Of all FDA-approved medications, around 34% target GPCRs. GPCRs are involved in blood pressure, heart rate, pain, mood, and immune response. In total, there are over 800 distinct subclasses of GPCRs in the human body. GPCRs are of extreme importance in medicine and are the most important protein receptors in the body.
When a drug attaches to a GPCR, the binding is just the first step to initiating a chain reaction of new chemical structures through the receptor, resulting in allosteric signal propagation. Simply binding is not an active GPCR signal. A drug that attaches to a GPCR without prompting the right changes is ineffective and may also inhibit the signal from being communicated.
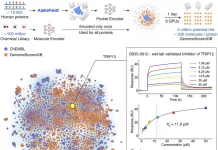
AlphaFold3, the cutting-edge model from Google DeepMind for structural biology, has gotten incredibly good at predicting how ligands may bind to and interact with proteins. It, however, cannot foresee the impact of such interactions and whether the protein in question remains inactive or is activated. This is where GPCRact comes in.
Teaching an AI to Think in Three Dimensions and in Process
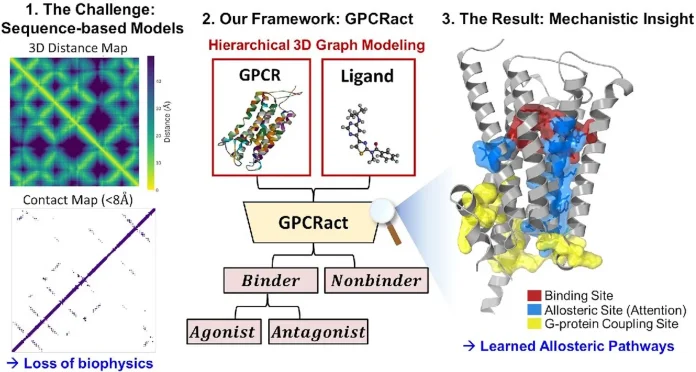
The main differentiating factor for GPCRact compared to other methods is how GPCRact views proteins. Most of the other models, like DeepREAL and AiGPro, view proteins as linear sequences of letters, where each ‘letter’ is an amino acid. It is reading just the name of a protein and not knowing anything about the protein itself. Son and Yi pointed out that this method has a serious limitation. For a given pair of amino acids in a sequence, the distance in sequence is a poor measure of how proximal the two amino acids are in a 3D representation. In this case, a positive correlation is present. Also, the sequence-based models have a clue of the evolutionary conservation, which is, in fact, a positive correlation with the least mobile areas of the structure that are going to be activated. So, these models focus more on the wrong areas of the protein that are the most critical.
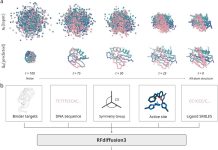
GPCRact completely disassociates with the sequence-first mentality. It constructs a graph from the actual 3D coordinates of the protein, with each residue and a few important sidechain atoms as nodes, and edges are drawn between atoms that are nearby in the 3D model of the protein. The ligand is viewed in the same way, as an atomic graph from its 3D conformer. This immediately solves the mismatch that earlier tools suffered from, where the ligand was modeled atomically but the entire protein was summarized to one residue-level code.
A Two-Stage Architecture That Mirrors Biology
GPCRact was built to replicate the natural process for which the model is intended. In Stage 1, the model centers on the binding event. A bidirectional cross-attention mechanism allows the protein’s binding site and the ligand to “communicate,” each modifying its internal representation based on the other. The result is a binding prediction and an “interaction signal” that describes the state of the binding site after the ligand has bound.
Stage 2 is where the real novelty lies. The interaction signal considers the full graph of the protein, not just the binding site or the allosteric site, where there are regions that physically move when a GPCR is activated. A local graph neural network first sends the signal to adjacent residues, and then a self-attention layer captures the signal on long-range spans across the whole structure. The ultimate result indicates whether the protein is poised towards an active or inactive conformation, and hence whether the drug is likely to be an agonist or an antagonist.
The model is also based on E(n)-Equivariant Graph Neural Networks, which means they can maintain the spatial underlying geometry in a protein structure, which is a very important property to keep the model’s comprehension of molecular architecture intact, independent of any arbitrary system of coordinates.
When the Hard Cases Are the Whole Point
Any model can look impressive on easy examples. What sets GPCRact apart is its response to genuinely difficult examples. Allosteric GPCRs with intricate signal propagation. The KAIST researchers determined that receptors with more than 200 allosteric residues that undergo significant repositioning between active/inactive states are ‘allosterically complex.’ Class C receptors are exemplary of this, particularly glutamate receptors.
GPCRact outperforms all contemporaries on standard GPCRs with sequence-based tracking models. On allosterically complex receptors, this gap becomes larger. AiGPro and DeepREAL, two of the current best models, drop 7-8 percentage points in these situations, while GPCRact drops barely 2 percentage points. This is a result of training of GPCRact on GPCRactDB, a novel database containing 200,000 GPCR interactions, the most any predictive tool has had to rely on, across 295 unique human GPCRs, a clear existing gap providing the base.
What Comes Next
While the GPCRact team continues work on discovering small molecules that act as agonists or antagonists, the team acknowledges that these two categories of pharmacological agents make up 95% of newly approved GPCR-targeting drugs, and that there are still options to explore, such as inverse agonists, partial agonists, and biased signaling. They also recognize the limitations of static protein structures and the need to integrate molecular dynamics to improve the modeling of protein flexibility.
The current implications of their work are already of great importance. A tool that goes beyond “does this molecule bind?” to ask “does it turn the receptor on or off, and to what downstream pathway?” fundamentally alters the nature of the computational drug discovery that can be done. It advances the field toward the much deeper level of mechanistic thinking that medicinal chemists use at the bench, while also striking the right level of transparency for researchers to validate the model’s reasoning against the current biological knowledge.
The GPCRactDB database and code are accessible on GitHub, so this is not just a proof of concept behind a paywall. It is something the wider community can develop further. In a field where so many lucrative AI technologies are evasive or proprietary, this is significant.
Article Source: Reference Paper | Reference Article | Code Availability: GitHub
Disclaimer:
The research discussed in this article was conducted and published by the authors of the referenced paper. CBIRT has no involvement in the research itself. This article is intended solely to raise awareness about recent developments and does not claim authorship or endorsement of the research.
Follow Us!
Learn More:
Anchal is a consulting scientific writing intern at CBIRT with a passion for bioinformatics and its miracles. She is pursuing an MTech in Bioinformatics from Delhi Technological University, Delhi. Through engaging prose, she invites readers to explore the captivating world of bioinformatics, showcasing its groundbreaking contributions to understanding the mysteries of life. Besides science, she enjoys reading and painting.