Picture a world where computers can not only translate languages but also decipher biology’s convoluted language. This is the exciting frontier of Large Language Models (LLMs) that could transform our knowledge of genes and cells, which are the foundation of all life forms. Researchers from the Center, Chinese Academy of Sciences, China, explore this intriguing crossroad. Genes, which are passed down from one generation to another, hold the truths of our being. Cells, the tiny factories that keep us alive, execute these instructions coded in genes. Decoding how genes and cells interact helps to unravel health complications, diseases, and even mysteries regarding evolution.
Traditionally, scientists have used gene sequencing to study these intricate associations. But LLMs present an alternative way forward with immense promise. These models are trained using huge volumes of text data, enabling them to understand complicated patterns as well as mappings between them. Perhaps scientists can get a breakthrough by passing such datasets through LLMs.
Do LLMs Understand Genes and Cells?
Chen Fang et al. carried out research to look at how well LLMs master the intricate process of genes and cells. On three main tasks, several leading LLMs were tested:
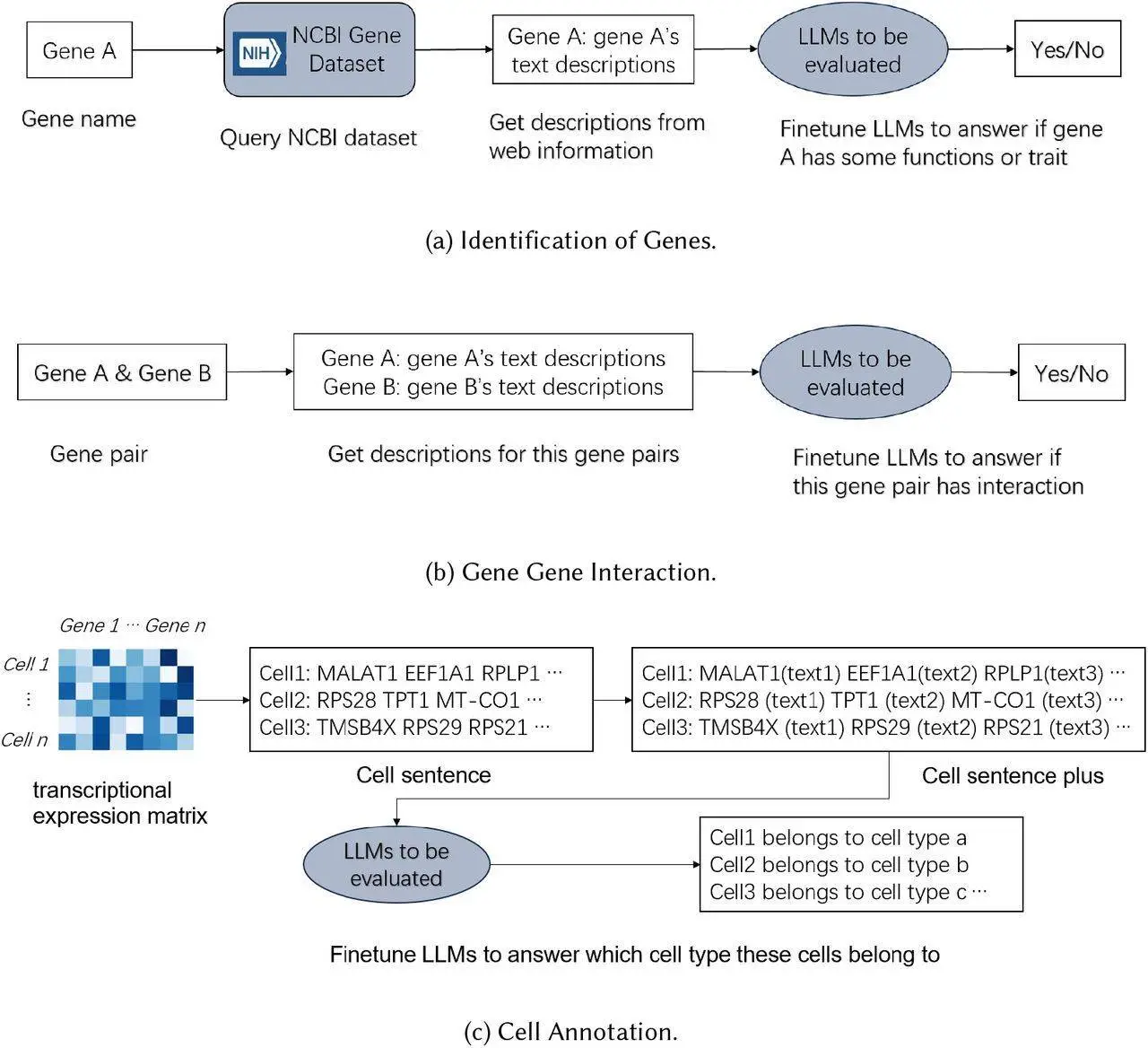
Gene Identification: Can LLMs zero in on individual genes within a flood of text?
Predicting Gene Interactions: Can LLMs foretell how various genes work synergistically or antagonistically?
Cell Annotation: Are the descriptions given by these LLMs enough to assign an accurate label to a cell?
The results were very positive. It was surprising that the LLMs performed the three tasks relatively well. This means they could successfully identify genes from scientific text, predict interactions between genes with some accuracy, and even annotate cells based on their gene expression profiles.
Going Beyond Names: A Better Way for LLMs to “See” Cells
One interesting aspect of this experiment had to do with how cells were presented to the model. Typically, scientists create what is known as “cell sentences,” which are simply lists of the most active genes in individual cells; however, such bare-bones explanations proved difficult for LLMs.
To solve this problem, a new approach called “cell sentence plus” has been developed by them. For LLM to analyze, this method involves adding brief functional descriptions to each of the gene names. That was a noteworthy success! In this case, cell sentence plus significantly increased the accuracy of cellular annotation tasks by giving more context.
LLMs: A Powerful Tool for Cell Biology
This study shows how LLMs are very important in the area of cell biology. These are some of the exciting possibilities:
- Faster Drug Discovery: LLMs would likely be able to analyze large datasets containing genes and identify potential drug targets at high efficiencies.
- Personalized Medicine: Through understanding the individual’s unique genetic structure, LLMs can assist in customizing medical treatments for maximum efficiency.
- Unraveling Complex Diseases: Since most diseases, such as cancer and Alzheimer’s, have intricate gene interactions, LLMs could play a role in decoding them.
Challenges and the Road Ahead
Even though we cannot deny the potentialities of LLMs, there are challenges ahead. They are trained on huge text datasets, and their data quality is critical. Additionally, scientific language can be complex and nuanced; hence, these subtleties may not be fully captured by present data sets. Furthermore, LLMs can not think critically and reason well. They can detect patterns in the data but may have trouble making sense of the underlying biology.
The Future of LLMs in Genomics: a Glimpse into a Transformed Landscape
However, LLMs hold greater value in genomics beyond addressing such challenges. Here is a sneak peek into what lies ahead as the technology matures and becomes seamlessly integrated with biological research.
- Unmasking Complex Diseases: For example, LLMs can go through huge datasets on people with complex illnesses like cancer, Alzheimer’s, or autoimmune diseases. By discovering subtle patterns in gene interactions that could be missed by traditional methods, LLMs may reveal new drug targets and treatment strategies for these diseases.
- Scaled Personalized Medicine: A time will come when doctors prescribe medical treatments that suit an individual’s specific genetic configuration using this technology. With this breakthrough, LLMs would help doctors analyze patients’ genomes to predict how they might react to various drugs; hence, it would pave the way for personalized medicine to be practiced at an unprecedented scale.
- Rapidly Automating Drug Discovery: Everyone knows that the drug discovery process is slow and expensive. LLMs have the potential to greatly speed up this process by enabling quick analysis of large molecule libraries that will help in the selection of those with some capacity to interact with distinct genes or cellular pathways.
- Synthetic Biology Grows Notably More Intelligent: The basic idea behind synthetic biology is designing and building biological systems for specific purposes. In particular, LLMs could be used to predict what kind of genetic changes would lead to functional consequences, thereby making it easier to “design” cells with certain properties.
- Enabling Genome Research for All: By using LLMs, cutting-edge genomics research can be made accessible to a larger community of scientists. Cloud-based LLM platforms and user-friendly interfaces would enable researchers who are not well-versed in bioinformatics to use LLMs in their studies.
The future of life itself will be fundamentally transformed by these applications and many others we cannot think about now.
LLMs may:
- Read the Non-Coding Genome: There might be hidden regulatory elements within those vast stretches of non-coding DNA earlier thought to be “junk.” Thus, LLMs should analyze these areas and establish their significance.
- Unravel the Quandaries of Evolution: By investigating numerous genomes, LLMs could provide additional insights into evolutionary mechanisms and the complex fabric of existence on Earth.
- Engineer the Future of Life: As our comprehension of genome broadens, LLMs could be significant in contributing to gene editing safe and ethical approaches as well as manipulating complex biological systems.
The future of LLMs in genomics is a story yet to be fully written. However, with further research and development, these mighty AI instruments might unlock a new period of discovery in life sciences that will shape our understanding of health, disease conditions, and what constitutes life itself.
So what do you think? Are we seeing the next big thing for cell biology with LLMs? Comment below or ask any questions! Let’s take this conversation forward and discuss how artificial intelligence may enlighten us on matters of life.
Article Source: Reference Paper | The code and data used in the study are available on GitHub
Important Note: bioRiv releases preprints that have not yet undergone peer review. As a result, it is important to note that these papers should not be considered conclusive evidence, nor should they be used to direct clinical practice or influence health-related behavior. It is also important to understand that the information presented in these papers is not yet considered established or confirmed.
Follow Us!
Learn More:
Anchal is a consulting scientific writing intern at CBIRT with a passion for bioinformatics and its miracles. She is pursuing an MTech in Bioinformatics from Delhi Technological University, Delhi. Through engaging prose, she invites readers to explore the captivating world of bioinformatics, showcasing its groundbreaking contributions to understanding the mysteries of life. Besides science, she enjoys reading and painting.