Imagine a teeming city where proteins are the movers and shakers. Each protein intertwines with others in partnerships that are vital to every cellular process. Nevertheless, unraveling these interactions, which are called protein-protein interactions (PPIs), forms the basis for both biology and drug development. Detecting PPIs has been akin to overhearing hushed conversations in a crowded room.
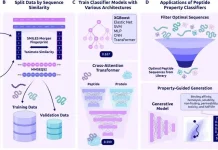
A recent study by Harvard Medical School researchers makes significant strides toward understanding the landscape of protein interaction. In this research, SPOC, a powerful classifier, was introduced, which improves the highly accurate identification of true PPIs compared to predictions made by AlphaFold-Multimer (AF-M), an innovative protein structure prediction tool. Predictions generated by the SPOC can be accessed and evaluated on the Predictomes database (predictomes.org). It serves as a repository for protein-protein interaction predictions, allowing users to explore and score their own predictions using SPOC.
Mapping Protein Interactions: A Challenge
Proteins are the basic units of life. They do many tasks, such as building organs and breaking down food. However, these proteins hardly work alone; most biological processes depend on complex collaborations between multiple proteins, and these interactions are very specific, like a lock and key. Understanding these PPIs is crucial in unveiling the mysteries of cellular machinery and developing new drugs for targeting particular interactions.
Scientifically speaking, yeast two-hybrid assays or co-immunoprecipitation have been traditionally employed to identify PPIs. However, these approaches are tedious, time-consuming, and often overlook important interactions.
Refinement is Necessary Due to AlphaFold-Multimer’s Rise
Over the past few years, AlphaFold-Multimer (AF-M) has been revolutionary in predicting protein structure. This state-of-the-art software employs deep learning to predict the 3D structures of protein complexes, thereby providing insights into PPIs. Notwithstanding its impressiveness, AF-M does suffer from one critical shortcoming: it can’t differentiate real from fake protein interactions well enough. Its predictions regularly contain a large number of false positives, thus making it hard to detect the genuinely significant interactions.
SPOC in Action: Unveiling New Interactions in Genome Maintenance
Picture yourself flying into the busy city of our cells, landing in a particular neighborhood that has the responsibility of providing security for our genetic material. This neighborhood is called genome maintenance machinery, and it includes many vital areas that are responsible for the repair of damages, replication accuracy, and ensuring DNA integrity. One way to prevent such diseases like cancer is through understanding how proteins within this area interact with each other.
This is where SPOC shines! For example, scientists used SPOC to look at relationships among nearly 300 human genome maintenance proteins or to analyze social network connections among 300 major players within this important cellular community! But it does not end there. Every protein can interact with any other one, making up a large number of interactions – almost 40000 protein pairs!
SPOC was able to sift through these voluminous documents like a super detective. It employed structural and biological aspects in the assessment of every pair’s likely interaction, enabling it to come up with high-confidence predictions. These high-confidence predictions provide exciting new directions for investigation.
Take a look at what SPOC discovered:
- Earlier unrecognized collaborations: SPOC identified previously unknown interactions among proteins that had not been known to interact before. The knowledge of possible new partners can provide vital insights into the functionality of genome maintenance machinery in general.
- Strengthening preexisting knowledge: Besides, it also confirmed established connections between genome maintenance proteins. This reinforces our current understanding of these important cellular activities.
- Revealing concealed relationships: A few highly confident predictions connected proteins belonging to different sub-processes within genome maintenance. These links indicate the possibility of crosstalk among these processes, thereby unveiling more intricacy in terms of interconnectivity within such a neighborhood.
But the most interesting thing? All these SPOC-predicted interactions do not lie idle in scientific journals. The researchers have made them accessible through predictomes.org, a friendly web interface. This helps scientists to:
- Explore the predicted interactome: View an interactive chart of human genome maintenance proteins and dive into the intricate network of their interactions.
- Focus on specific proteins: Look for interactions with a particular protein of interest and see what SPOC’s confidence score is for each prediction.
- Generate new hypotheses: Use this information to develop new concepts about how proteins within the genome maintenance machinery work together.
SPOC revolutionizes researchers studying genome maintenance by enabling them to identify highly reliable PPIs. This valuable tool reveals extensive dialogues that take place within cells, thus moving us towards a better understanding of how we preserve genetic integrity and potentially opening doors for new therapies against diseases caused by disruptions in these processes.
A User-Friendly Resource for Scientists
The findings of this study are a boon for researchers in various fields. The authors have made their data publicly available through a user-friendly web interface called predictomes.org. This website allows researchers to:
- Browse through the anticipated PPIs for SPOC-identified human genome maintenance proteins.
- Obtain the dataset for further analysis.
- Upload the protein sequences that are used in determining SPOC scores, which generally indicate how likely a given interaction is to have a true PPI.
The Future of Protein-Protein Interaction Mapping
Schmid et al. provide a foundation for the comprehension of protein interactions on a broad scale. SPOC allows scientists to identify actual PPIs from large-scale predictions. The well-curated dataset and user-friendly web interface further improve the accessibility and influence of this research.
With the development of more advanced protein structure prediction tools, like classifiers such as SPOC, we can expect significant progress in mapping out the intricate protein interaction networks that are within our cells. The vastness of this knowledge will certainly result in scientific breakthroughs inspiring discoveries in varied fields ranging from cause identification in diseases to focused therapeutics production.
Conclusion: A Brighter Future For Cellular Conversations
The crowded room where each person could only whisper to a few others represented the protein interaction landscape no more. Modern enhancements, such as SPOC, have given researchers incredible tools that enable them to overhear these important cellular conversations. Such an understanding of PPIs has huge consequences for future developments in biology and medicine.
Here are a few exciting possibilities on the horizon:
- Demystifying Disease Mechanisms: This will enable researchers to develop targeted therapies aimed at fixing the breakdown of specific protein interactions in diseases.
- Drug Discovery Revolution: By understanding how proteins interact intricately in particular pathways, we can come up with drugs that target those interactions precisely.
- Personalized Medicine: Based on their protein interaction network analysis, doctors might be able to come up with personalized treatment plans for complex diseases for patients.
Although still an ongoing process toward achieving a complete map of protein interactions, SPOC represents a major stride. As protein structure prediction tools advance and classifiers like SPOC get even better tuned, the cellular conversation will become more lucid. This new knowledge demonstrates a fresh era of scientific breakthroughs and transforms our understanding of health as well as medicine.
Ready to join the discussion? Check out the resources mentioned in this blog post, and keep an eye out for future developments in the fascinating field of protein-protein interactions!
Article Source: Reference Paper | The Predictomes database is available here
Important Note: bioRiv releases preprints that have not yet undergone peer review. As a result, it is important to note that these papers should not be considered conclusive evidence, nor should they be used to direct clinical practice or influence health-related behavior. It is also important to understand that the information presented in these papers is not yet considered established or confirmed.
Follow Us!
Learn More:
Anchal is a consulting scientific writing intern at CBIRT with a passion for bioinformatics and its miracles. She is pursuing an MTech in Bioinformatics from Delhi Technological University, Delhi. Through engaging prose, she invites readers to explore the captivating world of bioinformatics, showcasing its groundbreaking contributions to understanding the mysteries of life. Besides science, she enjoys reading and painting.