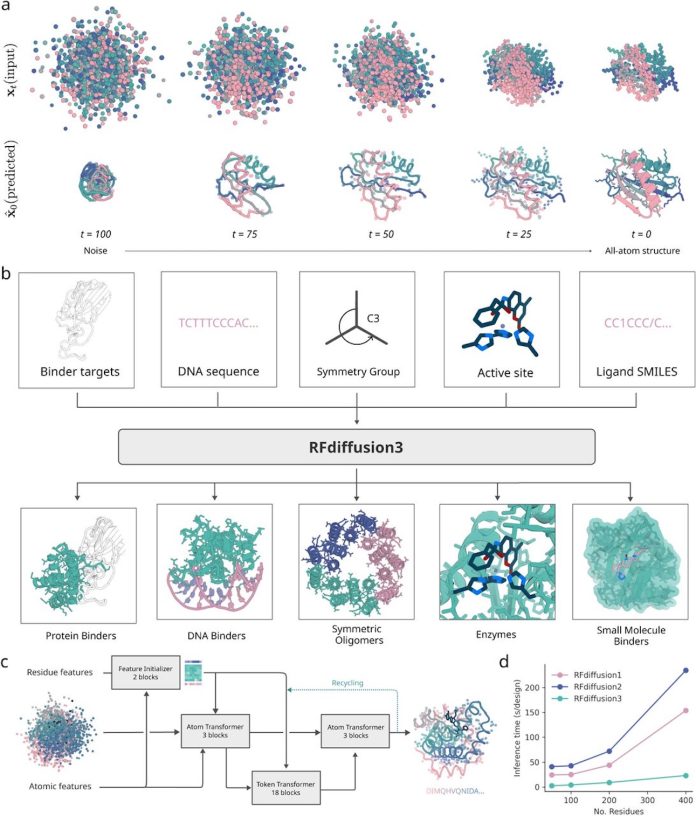
A major collaboration led by David Baker’s group at the Institute for Protein Design (UW) introduced RFdiffusion3 (RFD3), a next-generation version of the originally developed RFdiffusion. It is a generative diffusion model for general-purpose de novo protein design that works at the all-atom level, allowing precise modelling of hydrogen bonds, ligands, and nucleic acid interactions. It outperforms earlier RFdiffusion versions and, as part of the study, successfully designed and validated proteins that bind DNA sequences, created cysteine hydrolases, and improved the binding energy of small-molecule binders.
Key issues in Traditional De Novo protein designing
Most existing methods of de novo protein design generate proteins at the amino acid level (for example, RFdiffusion1 (year 2023) and RFdiffusion2 (year 2024) by the same team); good enough for designing monomers, assemblies, and even protein-protein binders. But they lack the accurate resolution for designing precise side chain interactions with non-protein atoms like ligands, small molecules, DNA/RNA binding proteins, etc.
It’s the side chains and individual atoms that form hydrogen bonds, salt bridges, van der Waals contacts, and π-π stacking. Residue or Backbone-only design doesn’t capture these fine details, which are the key concepts separating weak, strong, and site-specific binding, so only atom-level models can give us specificity, catalysis, and true biomolecular interaction design.
RFdiffusion: Architecture and Training: Challenges v/s the Innovation
To make all-atom generative protein design possible, researchers designed a transformer-based U-Net that encodes, processes, and predicts atomic coordinates with a special mechanism. Inspired by AlphaFold3’s diffusion module, it has three parts:
- Downsampling module: Encodes atomic and backbone-level features
- Sparse Transformer module: Processes token-wise information, reducing computational cost
- Unsampling module: Combines atomic and residue-level features
This creative design overcomes the challenge of coupling between both atom and residue level features; otherwise, side chains might not align properly with backbone geometry.
Instead of diffusing amino acid residues, RFD3 diffuses individual atoms, but the key challenge here was that, if the sequence is unknown (particularly in de novo design), the model doesn’t know in advance how many atoms each residue will have. This unpredictability makes it hard to represent proteins consistently in an atom-based framework.
To overcome this challenge, the authors standardized all residues to have 4 backbone atoms and 10 side chain atoms, padding smaller residues with virtual atoms. This trick makes atom-level diffusion far more practical for protein design.
Another major challenge of an all-atom generative model is its heavy computational effort. As highlighted by the authors, protein structure prediction differs a lot from protein design. One requires heavy modules and is computationally expensive (AlphaFold3), and the other does not need the same sequence-based feature extraction and can use a lighter conditioning.
The authors stripped down the conditioning module, making RFD3 much lighter and faster to train while still capable of handling atom-level complex constraints for designing tasks.
Insilico Results: Designing Some Proteins, Enzymes, and Small-Molecule Bindings
The authors demonstrated the performance of RFdiffusion3 by designing protein-protein binders, DNA binder proteins, and small molecule binders. RFD3 outperformed RFD1 on most therapeutic targets in protein-protein binder design, generating more unique and successful binding solutions.
For DNA binders, RFD3 jointly modeled protein and DNA conformations, with pass rates of ~8.7% (for monomeric) and ~6.7% (dimeric) designs with improved diversity.
When it comes to small molecule binders, RFD3 surpassed residue-level methods, generating complexes with rigid and flexible ligands.
Lastly, for enzyme design, RFD3 outperformed RFD2 on 90% of the benchmark active sites, especially in complex cases.
Performance and Benchmark Results Against Previous Models
In RFD2, side chain atoms were treated as a special new data type, while in RFD3. tip atoms are treated just like any other atom in the system. This makes the representation more uniform.
RFD2 could not even apply general atom-level constraints like hydrogen bond donor/acceptor, as it only represented a small subset of atoms. RFD3, on the other hand, allows direct conditioning of these properties.
The idea of representing residues with 14 atoms had been tried before in architectures similar to AlphaFold3’s Pairformer. But those were much more compute-heavy, unlike RFD3, which adopts this representation in a lighter and faster architecture.
Article Source: Reference Paper | Reference Article
Disclaimer:
The research discussed in this article was conducted and published by the authors of the referenced paper. CBIRT has no involvement in the research itself. This article is intended solely to raise awareness about recent developments and does not claim authorship or endorsement of the research.
Important Note: bioRxiv releases preprints that have not yet undergone peer review. As a result, it is important to note that these papers should not be considered conclusive evidence, nor should they be used to direct clinical practice or influence health-related behavior. It is also important to understand that the information presented in these papers is not yet considered established or confirmed.
Follow Us!
Learn More:
Saniya is a graduating Chemistry student at Amity University Mumbai with a strong interest in computational chemistry, cheminformatics, and AI/ML applications in healthcare. She aspires to pursue a career as a researcher, computational chemist, or AI/ML engineer. Through her writing, she aims to make complex scientific concepts accessible to a broad audience and support informed decision-making in healthcare.