New RNA transcripts have been discovered as a result of the growing amount of transcriptomic data. Bioinformatic tool developers have had difficulty separating long non-coding RNA (lncRNA) from protein-coding messenger RNA (mRNA). Currently, tools implementing deep learning architectures have been developed to accomplish this task, but their potential to discover sequence features and their interactions remain unexplored. The performance of deep learning tools was compared to others currently used in predicting lncRNA coding potential. In total, 15 tools were investigated, representing a wide range of methods. The tools were not only evaluated on known annotated transcripts but also on real-life data. Test sets with varying sizes and proportions of lncRNAs and mRNAs were used to assess the robustness and scalability of the tools.
Moreover, each tool was rated on its ease of use. In real-life datasets, deep learning tools performed well on most metrics and labeled transcripts similarly. All tools performed differently depending on the proportion of lncRNAs and mRNAs in the test sets. Top-ranking tools utilized computational resources differently, so the study’s nature may influence the decision to choose one over another. However, the results suggest that the novel deep learning tools outperform other tools currently used widely.
As the human genome contains large numbers of non-coding RNA transcripts, defining protein-coding transcripts from non-coding transcripts has become increasingly important. With the discovery of key long non-coding RNAs (lncRNA), advances in technology, and the completion of the human genome, all transcribed regions of the genome have been investigated, resulting in the discovery of many lncRNA molecules that are important to both healthy and diseased cells. A robust set of candidate non-coding transcripts is difficult to obtain based on sequence data due to many factors, including novel splice-site detection and transcript assembly. To address the question of coding potential and the distinction between protein-coding and long non-coding sequences, a variety of bioinformatics tools have been developed.
LncRNA detection tools primarily use sequence intrinsic features and/or statistics to distinguish coding from non-coding transcripts based on their length cutoff of 200 nucleotides (nt). In sequence analysis, the most commonly used features are open reading frame (ORF) length and Fickett’s score, calculated from sequence statistics such as nucleotide positions and K-mer composition. For protein-coding transcripts, 300 nt has been considered a minimum ORF length, and di- and trimers have been found to be more frequent in long non-coding transcripts than in protein-coding transcripts. Some mRNAs coding for short peptides has shorter ORFs than this cutoff, while some non-coding transcripts have longer ORFs.
Further, some tools analyze transcribed RNA sequences for homologies between known protein-coding genes and novel transcripts to help categorize them. Although this can be time-consuming, there are few sequence homologies to other non-coding RNAs for many of the known lncRNAs. Furthermore, some lncRNAs share homology with protein-coding genes and are thought to be derived from them. Hence, conservation of secondary structures has been proposed as being more common than sequence homology.
The list of features differentiating between protein-coding and long non-coding transcripts has been expanded in order to improve the classification of coding and non-coding RNAs. As a result, data becomes multi-dimensional. There is a limit to how many features can be included in traditional models, such as logistic regression. In order to train models using labeled training data, machine learning methods such as support vector machines (SVM) are typically used. However, the increasing complexity of these models can cause them to fail.
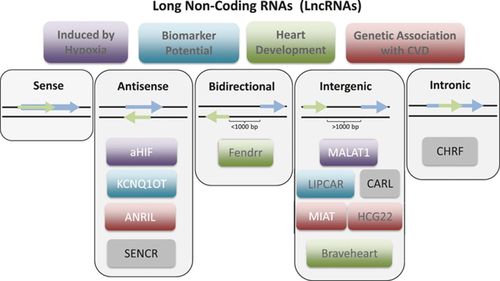
Image Source: https://doi.org/10.1161/CIRCRESAHA.114.304839
Based on the results of this study, LncADeep, mRNN, and RNAsamba performed better than most tools using other machine learning methods in nearly all datasets. In a real-life project, choosing the best deep learning tool may require balancing the computational and time resources. LncADeep may be chosen in many studies because accuracy weighs more heavily than computational resources. It may be preferable to use mRNN or RNAsamba online in other studies. It doesn’t matter what you choose, deep learning tools offer a fast and reliable way to identify lncRNAs, beyond what machine learning methods with only predetermined features can provide today since they can take into account features and feature interactions not included in current human knowledge. The only tool tested that was written in R was LncFinder. It is easy to install R packages, and many bioinformatics projects rely on R programs. Even though several other tools outperformed LncFinder in various performance metrics, LncFinder’s performance was quite good. LncFinder also requires a significant amount of computational memory during testing (64 Gbytes, Supplementary material), which may not be an option for some projects.
An evaluation of tools that implement deep learning methods was conducted against an evaluation of tools implementing other methods. The comparison of existing tools against deep learning tools did not take into account all existing tools. A variety of deep learning tools are available, such as lncRNA_MDeep and DeepCPP, which cover variations of the architectures evaluated here (DNN/DBN, CNN, and RNN). Zhang et al. performed in-house benchmarking on LncADeep, mRNN, and RNAsamba, concluding that LncADeep performs well. Furthermore, the testing confirms the findings of this study, that deep learning tools are more effective than tools based on other types of models.
Additionally, web-based user interfaces such as RNASamba and CPAT are helpful for adopting the best tools for wider systematic use in lncRNA prediction. In order not to give an advantage over tools that cannot be re-trained, none of the tested tools were retrained. The training on an earlier version of the human genome may have resulted in some tools performing better if trained on the current data. It is important, however, that a machine learning model be general and be able to work across a variety of datasets in order for it to be widely used. Therefore, these results may also reflect the generality of each tool.
Article Source: Reference Paper
Learn More:
Top Bioinformatics Books ↗
Learn more to get deeper insights into the field of bioinformatics.
Top Free Online Bioinformatics Courses ↗
Freely available courses to learn each and every aspect of bioinformatics.
Latest Bioinformatics Breakthroughs ↗
Stay updated with the latest discoveries in the field of bioinformatics.
Srishti Sharma is a consulting Scientific Content Writing Intern at CBIRT. She's currently pursuing M. Tech in Biotechnology from Jaypee Institute of Information Technology. Aspiring researcher, passionate and curious about exploring new scientific methods and scientific writing.





