Scientists perform the whole-transcriptome RNA sequencing and informatics of diabetes-induced sarcopenia model of db/db mice to illustrate the specific function of novel lncRNA Gm20743.
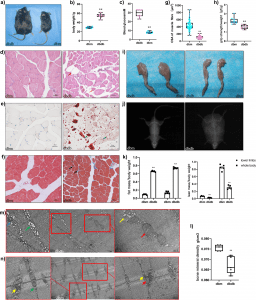
Image Source: https://doi.org/10.1038/s42003-022-03728-8
Type 2 diabetes mellitus raises the risk of sarcopenia, which is characterized by decreased muscle mass, strength, and function, although the specific process is yet unknown.
RNA whole-transcriptome sequencing and informatics were carried out on the db/db mouse model of diabetes-induced sarcopenia. In overexpression and knockdown-Gm20743 C2C12 cells, the detection of Mito-Sox, reactive oxygen species, Ethynyl-2′-deoxyuridine, and myosin heavy chain was carried out in order to ascertain the precise function of lncRNA Gm20743.
The main lncRNA-mRNA interactions were discovered using RNA-seq data and informatics, and this suggested that lncRNAs may play a regulatory role.
The scientists examined the potential roles of three key candidate lncRNAs, Gm20743, Gm35438, and 1700047G03Rik.
Furthermore, the findings revealed that lncRNA Gm20743 might have a role in controlling oxidative stress, mitochondrial function, cell proliferation, and myotube differentiation in skeletal muscle cells.
These discoveries considerably advance the knowledge of lncRNAs that may regulate muscle size, strength, and function in diabetes and serve as possible therapeutic targets for the sarcopenia brought on by diabetes.
The Unclear Mechanism of Diabetes-Associated Sarcopenia
Diabetes and obesity are the major health issues globally. Nearly 44% of type 2 diabetes mellitus cases are attributable to obesity, a global chronic health concern (T2DM).
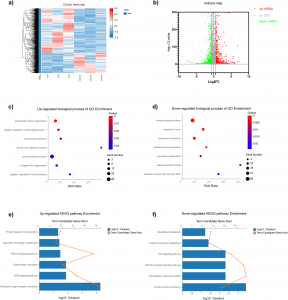
Image Source: https://doi.org/10.1038/s42003-022-03728-8
The skeletal muscle is the main organ for the metabolism of glucose. In addition to it, metabolic disorders, including diabetes and obesity, as well as insulin resistance, are all intimately associated with the loss of skeletal muscle.
A variety of pathological mechanisms, such as insulin resistance, oxidative stress, mitochondrial dysfunction, and inflammatory cytokines, are linked to the loss of skeletal muscle and intramuscular fat deposition.
Sarcopenia is a severe risk factor that could cause death and is a chronic consequence of diabetes that is made worse in patients with T2DM5. Loss of skeletal, muscular strength, mass, and physical function is a hallmark of sarcopenia.
Sarcopenia can emerge in midlife and is linked to several illnesses, including obesity and type 2 diabetes (T2DM), according to a new review published in The Lancet (2019).
T2DM patients were found to have decreased muscle mass, strength, and physical function when compared to healthy people.
Advanced glycation end-product buildup has also been linked to muscle atrophy in diabetic mice, according to a study conducted on rodents.
Sarcopenia linked to obesity and diabetes also worsens T2DM prognosis and lowers the quality of life due to increased risk of mortality, disability, and falls. The fundamental mechanism of diabetes-associated sarcopenia is still unknown, even though there are numerous identified risk factors and symptoms.
Role of LncRNAs
Primary transcripts are used to express the majority of the human genome. Although non-coding transcripts (also known as non-coding RNAs, or ncRNAs) make up about 98 percent of transcripts, only about 2 percent of transcripts (also known as messenger RNAs, or mRNAs) encode functional proteins.
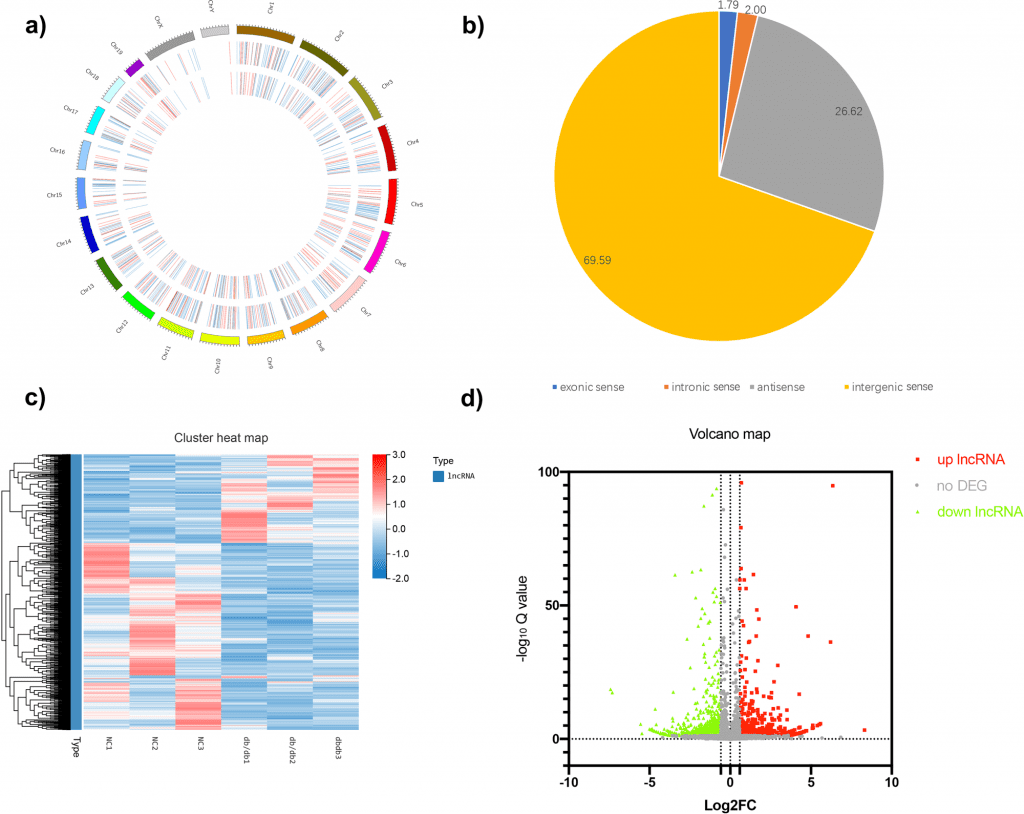
Image Source: https://doi.org/10.1038/s42003-022-03728-8
Long non-coding RNAs (lncRNAs) and microRNAs are the two primary subtypes of non-coding RNAs (ncRNAs).
LncRNAs are a subclass of longer than 200 nt RNAs that have little to no potential to code for proteins.
LncRNAs have greater tissue selectivity despite having less transcript quantity.
LncRNAs can bind to microRNA sites as competing endogenous RNAs, and the co-expression of lncRNA-mRNA in skeletal muscle cells plays a role in both pathological and physiological processes.
LncRNAs are possible therapeutic targets, biomarkers, or prognostic indicators that are closely associated with the development of diabetes and skeletal muscle.
The specific regulatory functions of lncRNAs during diabetes-induced sarcopenia are yet understood, as is the gene expression profiling of these molecules.
Demonstrations from the Study
In this investigation, the researchers used an acute stimulation in vitro model of palmitic acid (PA)-treatment in C2C12 myotubes along with an overt obese and diabetes db/db mice model in which muscle loss was evident.
Skeletal muscle fibers can be divided into slow (Type I) and fast-twitch (Type II) fibers. Patients with diabetes and muscle atrophy appear to be more likely to experience the preferred atrophy of type II fast-twitch fibers like the gastrocnemius (GAS).
The GAS from db/db mice and their littermates db/m mice were therefore subjected to whole-transcriptome sequencing.
After that, functional prediction and differential expression were carried out to investigate the gene-regulatory networks linked to diabetes-related sarcopenia.
Additionally, the candidate lncRNAs and their probable functions were discovered by co-expression network and informatics, including Gm20743 (log2FC = 1.57, q-value 0.001), and Gm35438 (log2FC = 1.90, q-value 0.001), and 1700047G03Rik (log2FC = 6.34, q-value 0.001).
The specific role of lncRNA Gm20743 in the onset of diabetes-related sarcopenia will be the focus of this investigation.
To learn more about how Gm20743 controls mitochondrial activity, oxidative stress, cell proliferation, and myotube differentiation, in vitro cultures of skeletal muscle cells with overexpressed and knockdown Gm20743 were used.
Together, these studies offer fresh perspectives on the molecular mechanisms governing the development of diabetes-related sarcopenia and suggest prospective targets for early detection and therapeutic applications.
Alterations in LncRNAs Regulation in Diabetes-Related Sarcopenia
According to mounting evidence, emerging lncRNAs have been linked to skeletal muscle growth and regeneration.
However, it is still unknown how lncRNAs regulate sarcopenia throughout the etiology of diabetes.
The given study demonstrates that full transcriptome RNA-seq in db/db mice identified thousands of differential gene expressions.
According to functional enrichment analysis, our findings showed that in db/db mice, pathways for “oxidative phosphorylation,” “metabolic pathways,” “muscle contraction,” and up-regulated “neuroactive ligand-receptor interaction” and “glucolipid metabolism” were considerably enriched.
Based on co-expression network analysis and functional prediction, they identified three key significant candidate lncRNAs, Gm20743, Gm35438, and 1700047G03Rik, as well as their probable functions. In db/db mice and PA-treated C2C12 cells, the putative lncRNAs and their closely linked gene expression were verified by RT-qPCR.
In this study, the scientists investigated the precise function of lncRNA Gm20743 in skeletal muscle cells. Their findings suggested that overexpressing Gm20743 could protect C2C12 cells from oxidative stress and mitochondrial dysfunction by lowering the level of intracellular and mitochondrial ROS in PA-treated cells.
The outcomes showed that Gm20743 overexpression might encourage myotube differentiation and cell proliferation in PA-induced C2C12 cells.
All of these findings point to previously understood relationships and novel alterations in the regulation of lncRNAs in diabetes-related sarcopenia, with a focus on the unique role of novel lncRNA Gm20743 in the onset of diabetes-related sarcopenia.
The Endpoint
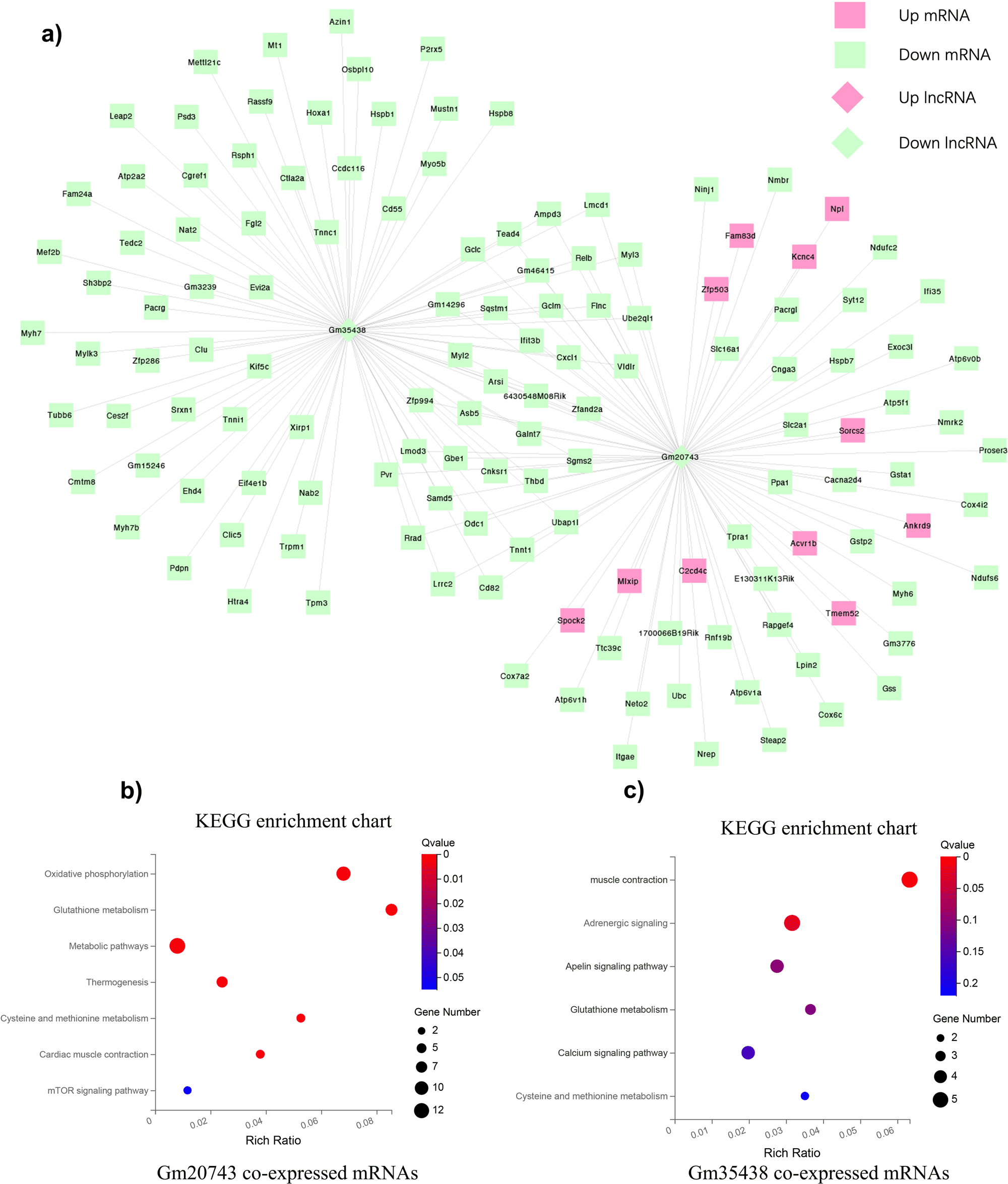
The scientists identified three candidate lncRNAs, Gm20743, Gm35438, and 1700047G03Rik, that may play crucial regulatory roles in diabetes-related sarcopenia based on lncRNA-mRNA co-expression networks.
Unexpectedly, the down-regulated lncRNA Gm20743 is an important lncRNA with 86 co-expressed differentially expressed genes (DEGs) in the co-expression network.
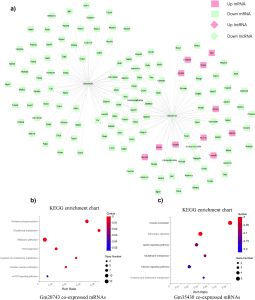
Image Source: https://doi.org/10.1038/s42003-022-03728-8
These highly correlated DEGs were primarily enriched in the down-regulated oxidative phosphorylation and glutathione pathway.
The energy released by food oxidation is transformed into ATP by mitochondrial oxidative phosphorylation.
According to studies, skeletal muscle mitochondrial function is harmed by ROS imbalance, oxidative stress, and impaired mitochondrial oxidative phosphorylation and ROS elimination systems.
Additionally, several studies have shown that the co-morbidity of obesity and diabetes causes mitochondrial dysfunction and activates oxidative stress, which ultimately results in a reduction in muscle mass and strength.
The findings on GSEA showed that the down-regulated oxidative phosphorylation pathway was highly enriched in db/db mice.
Additionally, TEM analysis of GAS tissue from db/db mice revealed altered mitochondrial morphology in comparison to the control group.
The performance of mitochondrial dysfunction in diabetic skeletal muscle was supported by these findings.
The researchers’ findings with the down-regulated glutathione route in db/db mice are consistent with a previous study that found the glutathione pathway has a beneficial major role in eliminating ROS and skeletal muscle mass by influencing redox homeostasis.
Additionally, the downregulation of significantly co-expressed genes (such as ATP1b1, Gclc, Gclm, and GPX4) that are connected to Gm20743 was consistent with the downregulation of the glutathione pathway and the oxidative phosphorylation pathway.
Therefore, in this investigation, they concentrated more on the specific role of lncRNA Gm20743 in mitochondrial function and oxidative stress.
Through in vitro tests, Lv-Gm20743 C2C12 cells revealed a lesser formation of mitochondrial and intracellular ROS in atrophic muscle cells treated with PA.
Mitochondrial complexes I and III produce ROS as a result of an imbalance in their enzymatic activity. It has been hypothesized that high levels of mitochondrial ROS cause downregulation of mitochondrial membrane potential and further decrease mitochondrial oxidative phosphorylation activity, which ultimately results in mitochondrial dysfunction. However, this study did not provide sufficient direct evidence to support this hypothesis.
These results imply that overexpression of Gm20743 ameliorates oxidative stress and mitochondrial function via reducing intracellular and mitochondrial ROS generation.
These results so clearly imply that overexpressed Gm20743 regulates the mitochondrial activity and redox equilibrium in PA-treated C2C12 cells via reducing intracellular and mitochondrial ROS levels.
The outcomes also show that overexpression of Gm20743 enhances myotube development and cell proliferation in PA-treated skeletal muscle cells.
It is significant to emphasize that GM20743 may influence mitochondrial activity and redox equilibrium to control skeletal muscle cell proliferation and myotube differentiation as a whole.
However, in public databases like GEO profiles and GTEx, the function of Gm20743 has not been identified in the skeletal muscle of any disease models.
This was the first time, to their knowledge, that the precise role of Gm20743 in mouse skeletal muscle cells had been described.
Additionally, they discovered that several human chromosome 1 sequences were extremely identical to the sequences of the mouse chromosome 1 lncRNA Gm20743 using gene alignment in the public human database, with the percentage of homologous genes reaching 70–80%.
Therefore, it is imperative to continue investigating the human gene homologous to Gm20743 as a target or biomarker for clinical diagnosis and therapy in the future. This work on the role of lncRNA Gm20743 in diabetes-related sarcopenia makes this possible.
Article Source: Yu, J., Loh, K., Yang, Hq. et al. The Whole-transcriptome Landscape of Diabetes-related Sarcopenia Reveals the Specific Function of Novel lncRNA Gm20743. Commun Biol 5, 774 (2022). https://doi.org/10.1038/s42003-022-03728-8
Learn More About Bioinformatics:
Top Bioinformatics Books ↗
Learn more to get deeper insights into the field of bioinformatics.
Top Free Online Bioinformatics Courses ↗
Freely available courses to learn each and every aspect of bioinformatics.
Latest Bioinformatics Breakthroughs ↗
Stay updated with the latest discoveries in the field of bioinformatics.
Tanveen Kaur is a consulting intern at CBIRT, currently, she's pursuing post-graduation in Biotechnology from Shoolini University, Himachal Pradesh. Her interests primarily lay in researching the new advancements in the world of biotechnology and bioinformatics, having a dream of being one of the best researchers.