We often underestimate the crucial role played by communications between cells to sustain living organisms. Flawless communication between cells is a necessity to maintain regenerative functions and to ensure the healthy development of tissues; even the slightest miscommunication can lead to severe consequences for the development of vital organs, therefore increasing vulnerability to diseases. Recent advances in genomics technology have allowed us to gain a deeper understanding of everything that occurs inside single cells. This has expanded our research opportunities to include studying the intricate relationships between cells within tissues and the elements that give distinct cell types their individual identities and functions. Researchers from Cambridge University and the University of Queensland collaborated to create CellPhoneDB v5, a bioinformatics toolset that provides a more thorough understanding of all these intricate relationships. To incorporate data from single-cell genomics, they combined statistical and computational methods with a repository consisting of a verified set of interactions between ligands and their corresponding receptors, all hand-curated by researchers. The multimeric essence that molecular complexes carry is captured by CellPhoneDB v5 as well.
What is cell-cell communication?
Communication between cells is facilitated through signals given by them to each other. There are three types of cell-to-cell communication: paracrine, endocrine, and juxtacrine signaling. Paracrine signaling involves the release of signaling molecules towards neighboring cells, after which they bind to them for activation. Endocrine signaling is carried out through hormones that are released from endocrine glands such as the hypothalamus in the brain or pituitary glands; these hormones transmit signals through the blood, interstitial fluids, or lymph fluids. In the third type of communication, juxtacrine signaling, the cells are in close contact with each other, where the ligand directly interacts with the receptor of the opposite cell.
To gain more clarity on how these signaling processes take place within living organisms, data from single-cell genomics has been modeled to understand the physiology of tissues in a holistic manner.
A basic overview of the updates in the latest version of CellPhoneDB
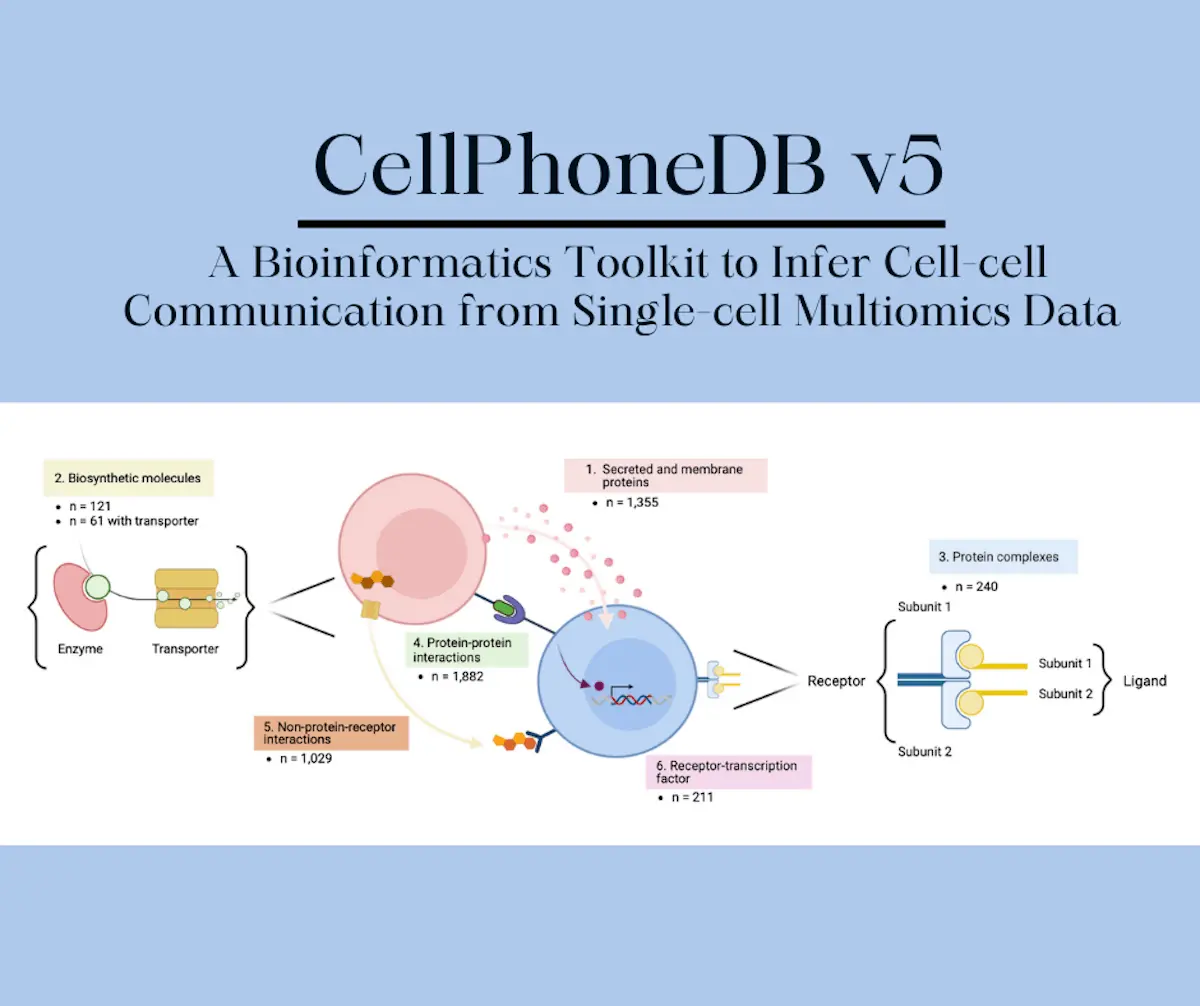
The fifth version of CellPhoneDB (v5) brings with it brand new additions that have increased the repository of the software by one-third of its previous volume through the incorporation of newly discovered interactions between cells that are moderated by ligands that are not proteins, for example, ligands that bind to G-protein-coupled receptors and endocrine hormones. G-proteins are the proteins that can bind to the nucleotides guanosine diphosphate and guanosine triphosphate.
The toolkit also employs a methodology based on differential expressions to deal with queries that are specifically based on interactions made with the user, along with enabling the use of novel computational methods that give greater importance to interactions between cells that are specific in nature. This feature uses different modes of single-cell activation, such as tissue-factor (TF) activation, modules like CellSign, or information related to the spatial orientation of molecules. The final cherry on top of this update is the module CellPhoneDBViz, which is utilized in visualizing the results for users in an interactive manner.
CellPhoneDB gives an estimate of the maximum number of ligands and receptors that can be expressed by a single cell. A combination of three essential components has been applied by the researchers to achieve this function in this update. The first component is a collection of reliable molecular interactions with determined roles in cell-cell communication. The second component takes the data to the level of tissues and incorporates transcriptomic data into the repository to give more context to all the interactions recorded in the dataset. The third and final component involves the application of approaches that interpret results that would normally be overwhelmingly complex in a simple visualization that is easy to understand which are sorted in order of relevance.
Important considerations by researchers when curating datasets
It is important to consider the scope and legitimacy of existing literature when studying cell-cell communications; regardless of which computational tools are used for generating inferences and no matter how powerful they may be, any incomplete or inaccurate piece of data related to cell-cell interactions will ultimately render all results useless in the end. Keeping this in mind, the single-cell genomics data used by the researchers contains peer-reviewed data from globally renowned databases such as UniProt and Reactome.
For two major reasons, the researchers also took the composition of the protein subunits into consideration to form an accurate representation of the interactions. The first reason is that some proteins are only functional if all the subunits are present; if even one of them is absent, the protein becomes non-functional. The second reason is due to the fact that ligands are very specific; protein complexes with multiple subunits can alter their own specificity to bind with their respective ligands. This change depends on the combination of heteromers that the complexes are associated with; the binding between them activates certain signaling pathways. Through the inclusion of heteromeric complexes in the database of CellPhoneDB, the number of false-positive results is reduced, increasing its reliability. The researchers also examined the last few steps in the biosynthetic pathways of interactions that are mediated by small molecules that do not present as proteins and included them in the dataset.
Applications of CellPhoneDB v5
This toolkit has already been applied in many key areas of medical research. It has been used to study how fibroblasts and epithelial cells present in the human endometrium interact when they come in contact with signaling hormones and to study how oocyte differentiation is driven through the compartmentalization of granulosa cells; both of these applications are relevant to studies pertaining to medical research in women’s reproductive health. Another application in the same area of research for CellPhoneDB is during the development of the embryo, where the interactions between maternal cells of the uterus and external trophoblast cells are evaluated. CellPhoneDB has been used in cardiac studies to recognize interactions between glial and trans-synaptic pacemaker cells located in the heart and neurons, respectively.
Conclusion
CellPhoneDB v5, according to the study’s experts, can shed light on the pathologies of many diseases and act as a roadmap for the creation of cutting-edge therapeutic strategies in in-vitro (or lab) environments. Modern computational techniques have been integrated into CellPhoneDB to assess cellular interactions, and the database has been improved. The code for the latest version of CellPhoneDB is available here.
Article Source: Reference Paper | CellphoneDB: Website
Important Note: arXiv releases preprints that have not yet undergone peer review. As a result, it is important to note that these papers should not be considered conclusive evidence, nor should they be used to direct clinical practice or influence health-related behavior. It is also important to understand that the information presented in these papers is not yet considered established or confirmed.
Learn More:
Swasti is a scientific writing intern at CBIRT with a passion for research and development. She is pursuing BTech in Biotechnology from Vellore Institute of Technology, Vellore. Her interests deeply lie in exploring the rapidly growing and integrated sectors of bioinformatics, cancer informatics, and computational biology, with a special emphasis on cancer biology and immunological studies. She aims to introduce and invest the readers of her articles to the exciting developments bioinformatics has to offer in biological research today.










[…] Unravel Cell-cell Communication with CellPhoneDB v5: A Cutting-edge Single-cell Multiomics Toolkit […]