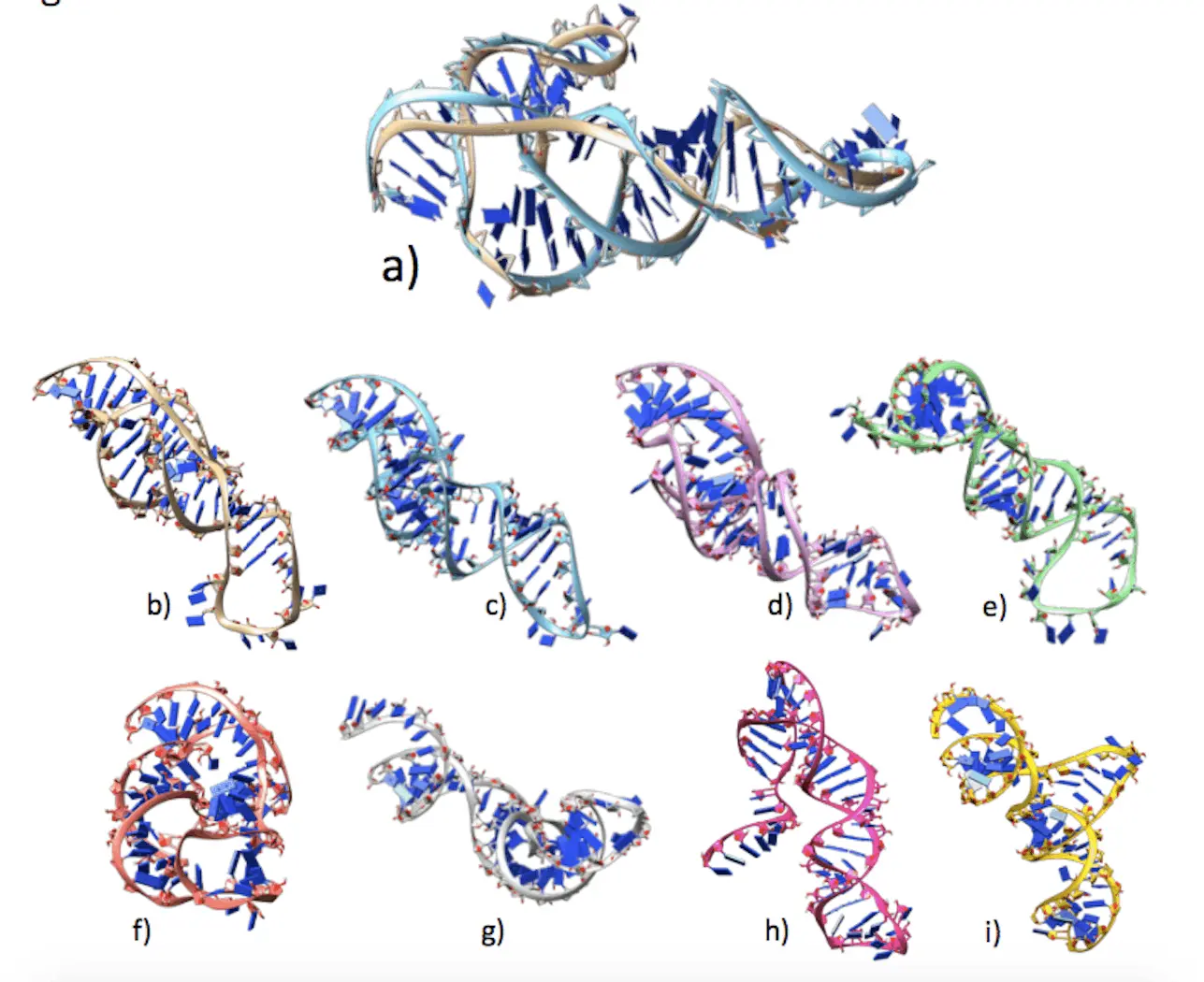
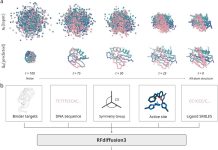
Before understanding RNA biology, one must grasp the three-dimensional structure of RNA. Because experimental procedures require a lot of labor and money, computer approaches have emerged. With the use of machine learning and physics-based concepts, these techniques can quickly anticipate RNA structures. On the other hand, their accuracy is still not as high as that of protein structure prediction. In this work, cutting-edge deep learning techniques for RNA structure prediction are benchmarked on a variety of datasets. The goal of this study by Nanyang Technological University researchers is to determine the variables that affect performance variation, including deep learning model architecture, RNA family variety, sequence length, RNA type, and multiple sequence alignment (MSA) quality. Deep learning model architecture, RNA family variety, length, type, and quality of multiple sequence alignment (MSA) all have an impact on RNA prediction techniques. While ML-based techniques are often more effective on most RNA targets, they do not work as well on unique or synthetic RNAs. The majority of approaches are unable to forecast non-Watson-Crick pairs, and the accuracy of MSA and secondary structure prediction is vital. The best prediction is made by DeepFoldRNA, with DRFold coming in second.
Challenges in Traditional RNA Structure Prediction
RNA molecules are studied by experimental techniques such as cryo-electron microscopy and X-ray crystallography, which are labor-intensive, costly, and time-consuming. These techniques are restricted to the study of a small subset of the enormous number of RNA molecules found in cells. Because RNA structures are dynamic and can adopt many conformations under varying physiological circumstances, they are more difficult to resolve. The 3D structure of RNA may now be predicted using computational methods, which combine machine learning, statistical analysis, physics principles, and available experimental data to produce realistic models of RNA structures quickly and affordably.
Entering Deep Learning: A Paradigm Shift
Computational methods have been developed for predicting RNA structures, either ab initio or template-based. Molecular dynamics is used in ab initio methods to simulate folding and find conformations with the lowest free energy. Fragment-based assembly, homology modeling, comparative modeling, and threading-based techniques are examples of template-based techniques. Some techniques combine modeling approaches based on fragments with coarse-grained folding. When compared to computational protein structure prediction, the accuracy of computational approaches for RNA structure prediction is still relatively low despite substantial advancements.
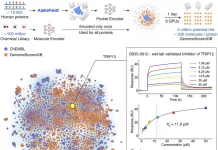
Protein structure prediction and RNA 3D structure prediction are similar, although RNA structures are thought to be more difficult to predict because there aren’t many experimentally established RNA structures in publicly accessible databases. About 1700 RNA-only structures are available in the Protein Data Bank, and most RNA families’ structures haven’t been experimentally established. The prediction procedure is further complicated by the fact that RNA structures include complex secondary structures and are more dynamic, subject to the effect of multiple factors. In recent years, deep-learning-based methods have shown promise in various bioinformatics tasks, including protein structure prediction. AlphaFold has revolutionized computational structural modeling, leading researchers to continue exploring these methods.
Assessing Deep Learning Approaches on Varied Datasets
This study assessed three datasets containing 66 target RNAs and seven RNA 3D structure prediction techniques. Two non-ML-based fragment assembly methods and five ML-based approaches were among the benchmarked methods. Performance evaluation took into account variables, including RNA kinds, dataset features, MSA, and RNA length. The performance of ML approaches was noticeably better than that of their non-ML counterparts. The most difficult dataset to work with was CASP15, which had a large number of synthetic or RNA-protein complexes. It’s possible that the abundance of targets disclosed before 2020 in the RNA-puzzles dataset contributed to its ease of usage. When modeling X-ray targets, the machine learning methods—which were mostly trained on crystallographic structures—showed superior accuracy. The recently assembled dataset, in which DeepFoldRNA has the lowest median residual standard (RMSD), offers the most realistic benchmarking set for assessing techniques in a blind prediction situation.
Findings of the Study
The length of the sequence and the kind of RNAs have an impact on the model quality of Natural RNAs. The best models are those of natural RNAs; the accuracy of models of RNA-protein complexes is mediocre. The hardest targets are synthetic ones. Longer RNAs have a lower TMscore, and as sequence length rises, the quality of projected models declines. However, when RNAs with sequence lengths shorter than 100 are taken into account, a positive association is seen between length and TMscore. Because longer RNAs are more difficult to anticipate in terms of base pairing, pseudo knots, and long-range interactions, their models are less accurate. The sequence length establishes the ideal range for RNA length.
The study discovered a weak relationship between the model quality and MSA for the RhoFold and RosettaFold2NA approaches, indicating that methods that merely accept MSA as input perform worse than those that accept MSA + secondary structure (ss) as input. The difference between the predictions made by RoseTTAFold2NA and RhoFold and those made by DeepFoldRNA and trRosettaRNA is more noticeable. RoseTTAFold2NA and RhoFold rely only on the MSA, which results in more inaccuracy in predicting base-pairing information and associated restrictions and lower TMscore models. DeepFoldRNA and trRosettaRNA leverage current methods for predicting the secondary structure.
The input ss determines the model quality of RNA structure prediction. Model quality was enhanced by using original PDBs ss rather than anticipated ss; however, for all but DRFold, the removal of input ss significantly decreased model quality. This emphasizes how crucial it is to use ss knowledge when predicting RNA structure.
Conclusion
This study shows that DeepFoldRNA and DRFold are the best models for RNA-based approaches, which have demonstrated the highest predictability for synthetic targets. When applied to artificial targets in the CASP15 dataset, these techniques perform poorly. Further development of methods should take into account the use of both secondary structure and MSA as input to enhance prediction. Improves in AlphaFold have been observed with metagenomic datasets, and the new RNA database can be used to create deeper MSAs for RNAs. Using a consensus ss predicted by many approaches, especially the most recent ML-based methods, is advised to improve input ss. The ML-based techniques will advance together with the number of RNA experimental structures solved and the RNA PDB database’s size.
Article Source: Reference Paper | Data availability: GitHub
Important Note: bioRxiv releases preprints that have not yet undergone peer review. As a result, it is important to note that these papers should not be considered conclusive evidence, nor should they be used to direct clinical practice or influence health-related behavior. It is also important to understand that the information presented in these papers is not yet considered established or confirmed.
Follow Us!
Learn More:
Deotima is a consulting scientific content writing intern at CBIRT. Currently she's pursuing Master's in Bioinformatics at Maulana Abul Kalam Azad University of Technology. As an emerging scientific writer, she is eager to apply her expertise in making intricate scientific concepts comprehensible to individuals from diverse backgrounds. Deotima harbors a particular passion for Structural Bioinformatics and Molecular Dynamics.