Universitas Gadjah Mada, Indonesia, scientists developed a low-cost, non-invasive method to rapidly sniff out COVID-19 by using an electronic nose named “GeNose C19”.
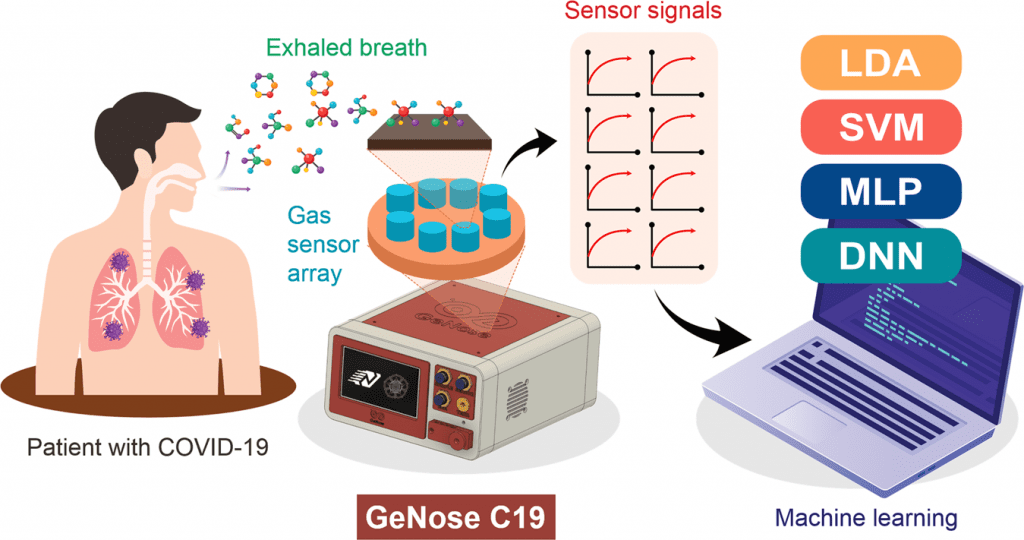
Image Source: https://doi.org/10.1038/s41746-022-00661-2
The method of RT-qPCR, which stands for a reverse transcription-quantitative polymerase chain reaction, has been widely utilized to identify the SARS-CoV-2 virus that causes COVID-19.
However, clinicians frequently prefer to use a combination of clinical signs and symptoms, laboratory tests, imaging measurements (such as a chest computed tomography scan), and multivariable clinical prediction models, including the electronic nose, to diagnose COVID-19 as opposed to using it alone.
Using a portable electronic nose (GeNose C19) that integrates a variety of metal oxide semiconductor gas sensors, improved feature extraction, and machine learning models, the scientists report on the invention and application of a low-cost, non-invasive technique to quickly sniff out COVID-19.
This method was tested using 615 breath samples total, including 333 positive and 282 negative samples, in profiling tests.
At two hospitals in Indonesia’s Special Region of Yogyakarta, the samples were taken from 43 positive and 40 negative COVID-19 patients, respectively, and validated with RT-qPCR.
To find the best pattern recognition techniques, four different machine learning algorithms—linear discriminant analysis, support vector machine, stacked multilayer perceptron, and deep neural network—were used.
Their findings imply that GeNose C19 is a highly promising breathalyzer for quick COVID-19 screening.
The COVID-19 Pandemic
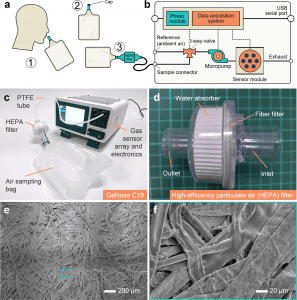
Image Source: https://doi.org/10.1038/s41746-022-00661-2
Infectious coronaviruses can cause respiratory and gastrointestinal infections in both people and animals.
Global health has faced significant challenges as a result of the discovery of a new coronavirus, formally known as SARS-CoV-2 (severe acute respiratory syndrome coronavirus 2).
The coronavirus illness 2019 (COVID-19), which is caused by the SARS-CoV-2 infection, was discovered in late 2019 in Wuhan, Hubei Province, China, and has since spread as the primary cause of a continuing and rising pandemic in more than 200 nations and territories worldwide.
SARS-CoV-2 has been regarded as the third most lethal pathogenic coronavirus over the past 20 years after the appearance of SARS-CoV in Guangdong Province, China, and the Middle East respiratory syndrome coronavirus (MERS-CoV) in Middle Eastern countries.
The other previously discovered human coronaviruses, such as HCoV-OC43, HCoV-NL63, HCoV-229E, and HKU1, only caused mild upper respiratory diseases in immunocompetent patients.
Significant mortality is linked to the COVID-19 pandemic, particularly in elderly and immunocompromised populations.
RT-qPCR as a Screening Tool
Since the pandemic’s start, the reverse transcription-quantitative polymerase chain reaction (RT-qPCR) approach has been regularly used to confirm the diagnosis of COVID-19.
The most extensively used test for SARS-CoV-2 detection to date is this diagnostic method that looks for the ribonucleic acid (RNA) of SARS-CoV-2.
Blood, sputum, feces, urine, bronchoalveolar lavage fluid, and nasal and pharyngeal swabs are a few examples of clinical samples that can be used.
A number of businesses and research facilities have created PCR-based detection kits that focus on at least two areas (genes) of the SARS-CoV-2 genome.
Compared to other diagnostic techniques, RT-qPCR is renowned for its excellent specificity (e.g., antibody and nucleocapsid protein antigen detection assays).
However, RT-qPCR still has a few drawbacks, including the requirement that it be carried out in a specialized laboratory, the requirement that invasive samples be done, and the high cost.
Due to their plentiful resources, affluent nations frequently do extensive RT-qPCR tests as a means of screening and epidemiological control.
However, similar qualities are regarded as opulent in developing nations, particularly those with middle-low incomes. Preventing pandemics requires ensuring that all nations have the capacity to conduct broad, quick screening followed by targeted isolation and control.
Many clinicians and researchers have attempted to use a combination of clinical signs and symptoms, laboratory tests, imaging measurements, and multivariable clinical prediction models, including the electronic nose, alongside RT-qPCR for validating clinical diagnosis results in order to get around the limitations of using RT-qPCR as a screening tool.
Because uncalibrated primer sets could potentially result in false-positive results for RT-qPCR, the detection procedure and primer sets frequently need to be first optimized.
In addition to RT-qPCR, more than 100 kits for COVID-19 diagnosis have been produced by various companies and are readily available on the market.
These kits use various working principles, such as isothermal amplification and lateral flow-based detection for nucleic acid targets, chest CT imaging, and immunoassays.
Need for an Artificial Olfactory System
Breath analysis offers an alternative as a quick and non-invasive detection method for sniffing out COVID-19 because it mostly affects the respiratory tract.
Patients, including those who are young and old, find it convenient and comfortable to undergo the procedure. Numerous metabolic pathways result in the hundreds of volatile organic compounds (VOCs) found in exhaled human breath.
Gas chromatography-mass spectrometry (GC-MS) and proton transfer reaction mass spectrometry (MS) could also be used to find them.
Acetone, methanol, ethanol, propanol, and isoprene are the VOCs that are most frequently found in exhaled human breath. Several additional substances are also found in exhaled breath. Some of them are benzene, acetonitrile, diallyl sulfide, allyl methyl sulfide, and diallyl disulfide.
In general, a number of VOCs can be used as non-invasive biomarkers for a variety of respiratory disorders, such as esophageal and gastric cancers, lung cancer, asthma, rhinovirus-induced wheezing, influenza infection in swine, and tuberculosis.
According to recent research, COVID-19 could also be identified utilizing a breath analysis method based on VOCs employing GC-IMS (near patient GC-IMS).
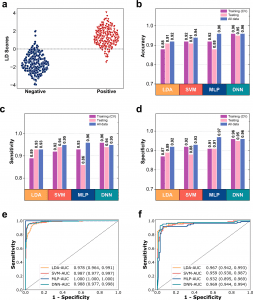
Image Source: https://doi.org/10.1038/s41746-022-00661-2
The MS analysis is still time-consuming, expensive, and difficult to apply for bedside clinical practice despite its excellent accuracy, capacity to distinguish VOC types properly, and utility in pathophysiological research.
Additionally, this technique alone is neither practical or practicable for rapid COVID-19 testing of a large human population, especially if the target detection time should only be a few minutes.
In order to assess VOC mixes in exhaled breaths, an electronic nose—an artificial olfactory system made up of a variety of integrated gas sensors with various active layers and artificial intelligence (AI) to differentiate complicated odors—can be used.
The sensors react uniquely to the different VOC percentages while they are exposed to the breath. Each odor, which represents a different VOC combination, can then produce a specific sensor signal pattern.
Such a VOC pattern is referred to as a breath-print for exhaled air. Therefore, complex VOC mixes can be differentiated from and categorized at high throughput without having to identify individual molecule components utilizing pattern recognition methods, including machine learning. An electronic nose is fast (producing results in a matter of minutes), portable, reasonably priced, and simple to use.
Exhaled breath analysis utilizing an electronic nose is appealing since it is non-invasive, quick, safe, and simple to use from the standpoint of patients in the case of COVID-19. The electronic nose has recently been employed as a pre-operative screening for COVID-19.
Previously, it was used in clinical settings to diagnose and monitor respiratory and urologic disorders (such as ventilator-associated pneumonia, TB, and kidney failure).
Following the ongoing evolution of COVID-19 within patients’ respiratory systems, VOC profiles are anticipated to vary. The SARS-CoV-2 and its particular interaction with the host are responsible for their complicated modification.
The Electronic Nose: The Scientists’ View
Several thin-film and micro-/nanoscale gas sensors can be used as the primary detecting elements in the electronic nose, depending on their operating principles and constituent materials. These include gravimetric sensors, chemoresistive sensors, and colorimetric sensors.
Due to their good qualities (i.e., low cost, quick response time, straightforward measurement setup, high durability, and extended lifetime), chemoresistive metal oxide semiconductor gas sensors have proven to be popular, particularly for VOC detection.
When many active material types are utilized simultaneously, the disadvantage of such sensors with regard to the moderate selectivity can be resolved.
Then, using AI-based data post-processing, their distinctive signals can be found and integrated. Consequently, the electronic nose, which consists of a number of chemoresistive metal oxide semiconductor sensors, has been utilized in several domains of environmental monitoring.
Another study has evaluated the evidence supporting the possible use of electronic nose technologies to identify and examine the COVID-19 breath pattern signature produced by the VOC-based target biomarkers.
In order to simulate the potential detection of COVID-19, an electronic nose made up of commercial metal oxide semiconductor-based sensors (i.e., MQ-135 and MQ-2) that can detect gas concentrations ppm of carbon monoxide, acetone, and alcohol was employed. The National Aeronautics and Space Administration of the United States had filed a patent for the suggested technology as a portable metabolic analysis device.
Diagnostic electronic nose systems were recommended to include a variety of metal oxide semiconductor-based gas and electrochemical sensors as the infectious disease got better in order to enable accurate monitoring of exhaled breath biomarkers of disease and modifications connected to the disease pathogenesis.
According to the authors, no other published study has yet looked into the possibility of using electronic noses to identify COVID-19 patients in a clinical bedside scenario.
Due to this, the scientists created the GeNose C19 portable breathalyzers in this study by combining a variety of metal oxide semiconductor gas sensors, machine learning analysis, and a breath sample setup.
To verify the system’s capacity for identifying differences between the exhaled breath patterns of the two subject groups, it was used in clinical experiments at two hospitals in Indonesia. In order to determine the produced device’s greatest possible accuracy, four different machine learning algorithms were tested.
The Endpoint
This study was designed to provide proof-of-concept evidence that COVID-19 infection can be predicted using a breath sample and detection.
The estimated performance values in our study essentially demonstrate the DNN algorithm’s dependability in estimating the training and testing data of breath samples, indicating the tremendous potential of the GeNose technology, strengthened by the DNN algorithm, to be employed as a COVID-19 screening tool.
Here, the scientists used an “open-label” study design, meaning that before undertaking sampling and dividing the sampled data into case and control groups, they already knew the individuals’ COVID-19 status.
They built their initial AI database using the breath sample pattern profiles from each respected group as training data. All of the data were validated using the RT-qPCR test results and clinical data. To find out more about the characteristic VOCs for COVID-19, a combined measurement of GeNose C19 with GC-MS will be done soon.
A study on the clinical diagnosis of COVID-19 with a greater number of exhaled breath samples is currently being conducted to demonstrate the potential of GeNose C19 as a quick COVID-19 screening tool using a cross-sectional design and double-blind randomized sampling. This is an important step in the system development process.
Article Source: Nurputra, D.K., Kusumaatmaja, A., Hakim, M.S. et al. Fast and noninvasive electronic nose for sniffing out COVID-19 based on exhaled breath-print recognition. npj Digit. Med. 5, 115 (2022). https://doi.org/10.1038/s41746-022-00661-2
Learn More:
Top Bioinformatics Books ↗
Learn more to get deeper insights into the field of bioinformatics.
Top Free Online Bioinformatics Courses ↗
Freely available courses to learn each and every aspect of bioinformatics.
Latest Bioinformatics Breakthroughs ↗
Stay updated with the latest discoveries in the field of bioinformatics.
Tanveen Kaur is a consulting intern at CBIRT, currently, she's pursuing post-graduation in Biotechnology from Shoolini University, Himachal Pradesh. Her interests primarily lay in researching the new advancements in the world of biotechnology and bioinformatics, having a dream of being one of the best researchers.












