A research team led by Prof. Nir Hacohen from the Broad Institute of MIT and Harvard, Cambridge, designed the study Dictionary of Immune Responses to Cytokines at single-cell resolution. This dictionary is a compilation of single-cell transcriptomic profiles of over 17 immune cell types in response to each of 86 cytokines (>1,400 cytokine–cell type pairings). Immune Dictionary expands our understanding of the activation states of all immune cell types, generates new hypotheses for cytokine functions, highlights the pleiotropic effects of cytokines, and offers a framework for determining the roles of particular cytokines and cell-cell communication networks in any immune response.
Purpose of Immune Dictionary
A large class of tiny, secreted proteins known as cytokines attach to specific receptors on target cells to initiate local or systemic action. This initiates downstream signaling and coordinates immune system cell types’ actions. Numerous illnesses, such as cancer and autoimmune diseases, are treated with cytokine-based medicines and cytokine antagonists. A large number of studies have been conducted depicting the central role of cytokines in immune function. However, a generalized global view of the cellular responses of each immune cell type to each cytokine was lacking. To fix this breach, the Immune Dictionary was introduced.
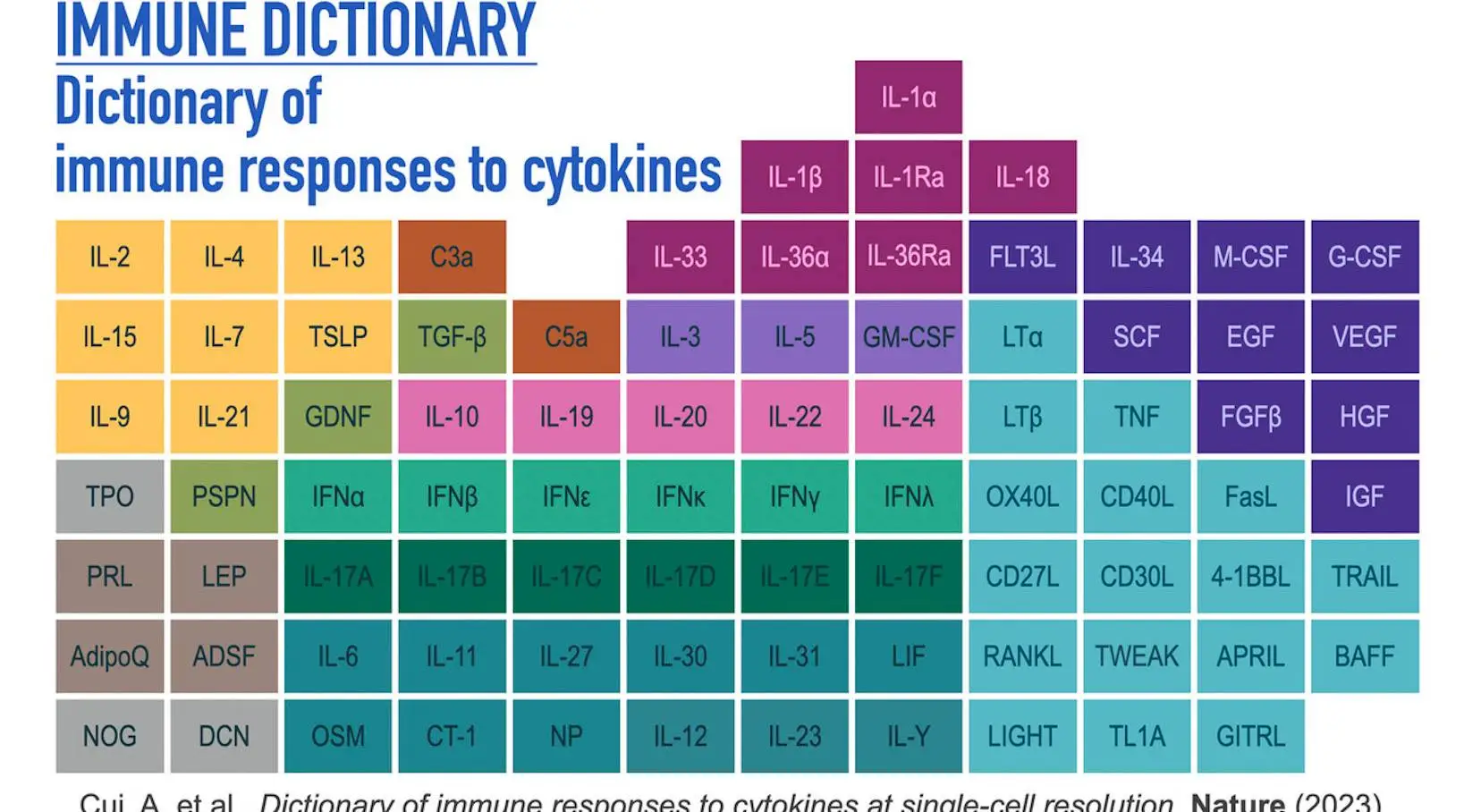
In order to create a large-scale perturbational single-cell RNA sequencing (scRNA-seq) dataset of the immune system, the researchers methodically profiled single-cell transcriptomic (single-cell RNA sequencing, or scRNA-seq) responses to 86 cytokines across more than 17 immune cell types in mouse lymph nodes in vivo. This allowed them to obtain a comprehensive view of cellular responses to cytokines. The 86 cytokines comprise representative molecules from other families with cytokine functions as well as the majority of members of its major families, such as interleukin-1 (IL-1), common γ-chain/IL-13/thymic stromal lymphopoietin (TSLP), common β-chain, IL-6/IL-12, IL-10, IL-17, interferon (types I, II, and III), complement, and growth factor families.
Unveiling the Immune Dictionary
Cell-type-specific cytokine responses
The researchers examined how various cell types react to the same cytokine by doing a cytokine-centric study of this lexicon. The study showed that numerous cell types are impacted by cytokines, which specifically alter gene expression. Examples of these include TNF, IL-1β, and IFNβ. While some cytokines affect a small fraction of cell types, most upregulated genes are specific to one kind of cell. The DEGs that are most frequently shared are IL-4, IL-2, IL-15, IL-18, IFNα1, and IFNβ.
IFNα1, IFNβ, IL-1α, and IL-1β are among the cytokines that compactly produce complicated cytokine responses in a variety of cell types. In response to cytokines, all of these cytokines are elevated collectively, along with co-expressed genes. While IL-1α and IL-1β generate highly cell-type-specific GPs (expressed genes) with distinct biological processes, IFNα1 and IFNβ elicit common antiviral GPs across practically all cell types. These GPs augment the known activities of multiple cell types: Treg cells induce Hif1a and Ctla4, MigDCs and Langerhans cells upregulate migratory programs, and neutrophils upregulate chemokine and inflammatory genes.
Cytokine-driven cell polarization states
In order to determine the cell states brought on by cytokines, researchers subsequently carried out a cell-type-centric study of the dictionary. Major immune cell polarization drivers include cytokines, such as certain ones that cause macrophages to adopt pro-inflammatory M1-like or reparative M2-like states.
The research finds that 66 main polarization states—including IFNγ, IL-4, and IL-13—are brought on by cytokines. These states exhibit important biological programs and are enriched for cells treated with cytokines. When exposed to IL-1α, IL-1β, or IL-36α, monocytes express more Il1b, which could lead to the development of an inflammatory feedback loop. Other polarization states are triggered by IFNα1, IFNβ, and TNF, which makes previously known polarization states in immune cell types visible. While NK cells are well-known for their cytotoxic properties, other uses are continually being found for them. IL-1α and IL-1β generate a unique ‘NK-c’ state that has little association with other NK cell states and does not have an overexpression of cytotoxic molecules, whereas IL-2, IL-12, and IL-15 induce a state with elevated production of cytotoxic genes. More than 1,000 genes are increased when NK cells are activated by IL-18, primarily in the ‘NK-f’ state. This suggests that the NK cell–IL-18 axis has multiple roles in the immune system. Cytokines polarize B cells to express different GPs and generate different polarization states in T cells.
Researchers discovered that Langerhans cells and MigDCs shared cytokine drivers and polarization states. TNF may have improved antigen presentation and T-cell interaction by upregulating the levels of Cd40 and Ccl22. DC migration was impacted by the overexpression of Nr4a3, which was brought about by GM-CSF and IL-1 family cytokines, as well as an increase in Ccr7 expression. This demonstrates the adaptability of NK cells as well as all other immune cell types.
Cytokine production–response map
In networks of immunological cell-cell communication, cell type and sources of cytokine production are important factors. While ILCs and basophils, two other uncommon cell types, also express a significant number of different cytokines, FRCs express the greatest number. The abundance of a certain type of immune cell and the production of cytokines are inversely correlated. Cell-cell interactomes that map out the pathways via which cells in the immune system can communicate demonstrate that FRCs can affect almost all cell types by producing a wide range of cytokines. High interconnectedness in the immune system is demonstrated by the fact that virtually all cell types can influence almost all other cell types through at least one cytokine.
Treatment with cytokines modifies receptor expression, which may make cells more susceptible to additional cytokines. TNF and IFNβ cause cDC1 cells to overexpress Cd40, which impacts T, NK, and DCs. Certain cytokines, such as IL-1α and IL-1β, elicit reactions even in the absence of highly expressed receptors, exposing a variety of cell-to-cell relationships.
Immune Response Enrichment Analysis
Despite transcriptome data being essential for researching immune responses and illnesses, it frequently lacks details on the stimuli that lead to the observed cell states and their purposes. Researchers have created Immune Response Enrichment Analysis (IREA), a software that evaluates the enrichment of cell polarization or cytokine signatures in transcriptomes to infer cytokine responses and cell-cell communication networks. A single-cell transcriptome dataset from immune cells in mice’s tumors receiving anti-PD-1 checkpoint-blocking medication was subjected to IREA analysis. IL-2, IL-12, and IL-15-induced polarization of other cell types, such as NK cells, into a cytotoxic ‘NK-e’ state was discovered by IREA. In cells treated with anti-PD-1, the immunosuppressive cytokine TGFβ1 exhibited the most negative reaction, suggesting the involvement of a cytokine network in the effective response to immunotherapy. Applying IREA to a severe case of COVID-19 infection exposed B and T cells to cytokine responses, which were consistent with previously reported elevations in plasma cytokines that control lymphocytes.
Future Directions
This immune dictionary can provide researchers with opportunities to study different cytokine doses, time, biological contexts, and combinations of stimuli.
Methods Implemented
This study developed a comprehensive atlas of immune cell transcriptional responses to a panel of 86 cytokines. The researchers injected each cytokine subcutaneously into mice, collected draining lymph nodes, and performed single-cell RNA sequencing. They analyzed differential gene expression, gene programs, polarization states, cytokine production, and cytokine receptor expression across diverse immune cell types in response to cytokine stimulation. Furthermore, they developed a computational platform called the Immune Dictionary for interactive exploration of the data and tools, enabling users to map their data onto this reference database to gain biological insights. This rich dataset and analytical framework offer an invaluable resource for elucidating immune cell communication pathways and functions.
Conclusion
The study shows the complexity and adaptability of immune cells by offering a high-resolution picture of the cytokine network. It demonstrates how different reactions in every type of cell can be triggered by a single cytokine, resulting in a coordinated multicellular immune response. Each immune cell type’s cytokine-induced polarization states are also identified in the study, demonstrating the immune cells’ malleable reactions to external stimuli. Reanalyzing responses in any subpopulation of cells is made simple by the dictionary.
Article Source: Reference Paper | Reference Article | IREA is available at the Website
Learn More:
Deotima is a consulting scientific content writing intern at CBIRT. Currently she's pursuing Master's in Bioinformatics at Maulana Abul Kalam Azad University of Technology. As an emerging scientific writer, she is eager to apply her expertise in making intricate scientific concepts comprehensible to individuals from diverse backgrounds. Deotima harbors a particular passion for Structural Bioinformatics and Molecular Dynamics.
















[…] Broad Institute Researchers Unveil a New “Dictionary” of Immune Responses to Cytokines a… […]