Investigators from Brigham and Women’s Hospital developed a novel approach to identify time-frequency peaks from sleep brain waves revealing heterogeneity between both healthy and neurologically diseased subjects. The approach provides fingerprint identification unique to each individual and identifies the potential for neurodegenerative disease phenotyping and biomarker identification.
Introduction
Traditionally, the identification of brainwaves in the form of oscillatory activity using the electroencephalogram (EEG) was used to determine the neural activity and responses across the time frame. With continuous oscillatory components, these brainwaves also present a short burst of activity called transient oscillations. Since these bursts are often short-observed, it appears difficult to draw inferences. The sleep spindles EEG hallmark defines the NREM (non-rapid eye movement) stages that are heavily associated with sleep-related functions, neurodevelopment, aging, and neurological disorders such as schizophrenia and Alzheimer’s disease.
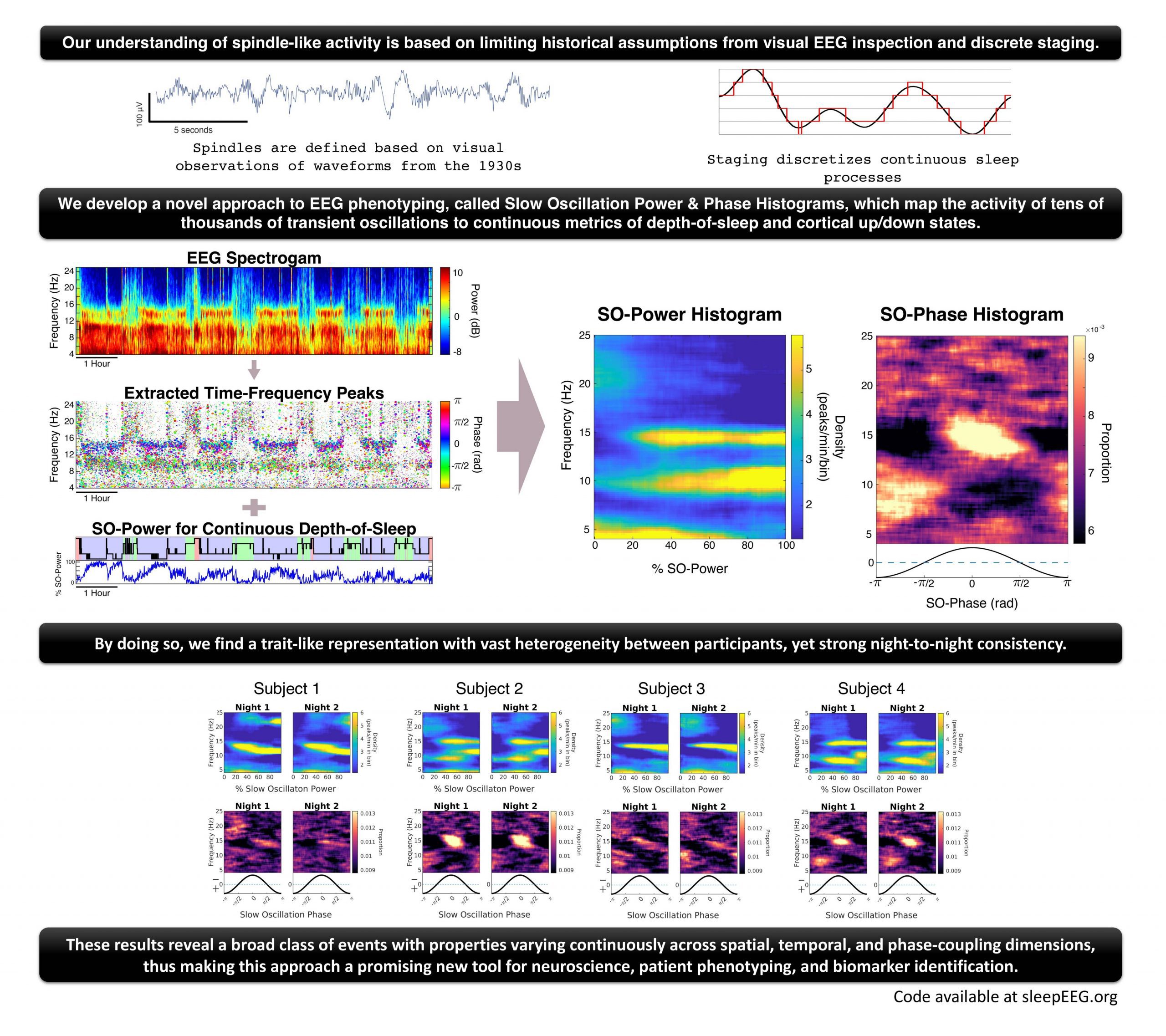
A team of researchers from Brigham and Women’s Hospital developed a novel approach to improve the understanding of these sleep spindles to generate answers. The approach for advanced EEG phenotyping is also called Slow oscillation power or SO-power (continuous metrics of depth-of-sleep) and Phase histograms (a correlate of cortical up/down states). The team has utilized a two-fold power phase representation providing robust phenotyping among the individuals. The transient oscillations were leveraged as 3D peaks in time-frequency domain topography, and the confined peaks as TF-peaks (Time-Frequency peaks) were observed as spindle blobs in the spectrogram, which were associated with defining the functioning of oscillatory events.
In order to analyze the TF-peaks within a given state of the brain, the researchers characterized a sleep continuum microarchitecture which was clinically inspected using standards and stages such as a wake, rapid-eye movement, and non-rapid eye movement. Using SO-power due to its characteristic of simple usage and objectivity also is particularly useful in continuous characterization. By combining the base concepts for TF-peaks, distributional analysis, and benchmarking continuous sleep dynamics, a novel approach was developed for Electroencephalogram phenotyping.
The approach adopted for the development included the following:
1) Identifying the TF-peaks in the sleep EEG spectrogram across both time and frequency.
2) Represent continuous standards and short-scale neural dynamics by extracting SO-power and SO-phase.
3) Characterizing the TF-peak rate as a function of metrics such as SO-power, SO-phase, and TF-peak oscillation frequency.
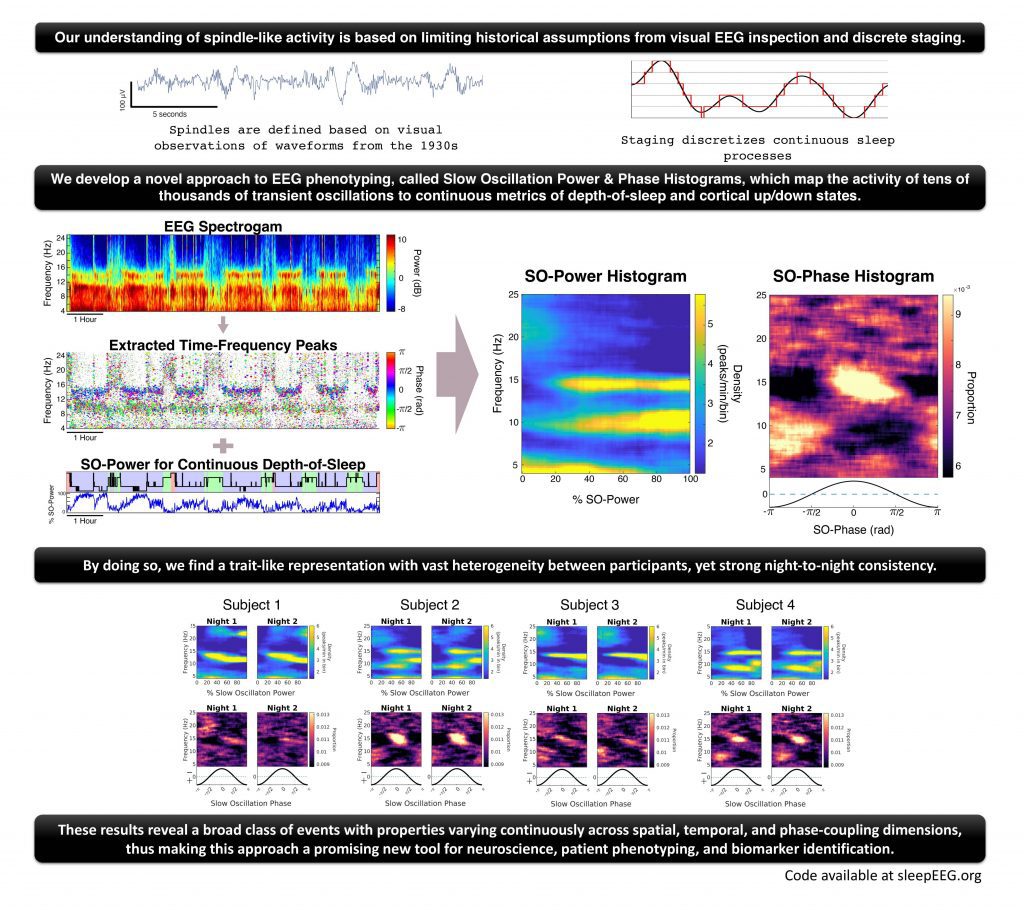
Image Source: 10.1093/sleep/zsac223
The methodology consisted of experimental data obtained for 17 healthy individuals put as controls (HC) and a total of 21 schizophrenia patients. The data was obtained from an earlier study conducted by Wamsley et al. in 2012. The participants were encountered with continued polysomnography for two nights, and EEG was digitally obtained by the team. Within the multi-paper spectrogram results, heterogeneity within the healthy population was distinctly observed, and the graphs were consistent throughout the duration. The observation led to personalized characterization within individuals who could potentially be used to identify different phenotypic traits of sleep EEF. Upon expanding the study dimensions for sleep continuum and sleep-depth coupling, the variations were addressed.
Considering sleep as a continuous process with time variations team characterized TF-peak and phase coupling dynamics to extract the complex brain temporal activity during sleep. Across the results and observations, the SO-power spectrogram showed an improved depiction of TF-peak activity in terms of understanding the sleep continuum. The comprehensive approach also showed an increase in dimensionality as compared to the classic analysis.
The methodology was used to characterize the TF-peaks, the team determined the SO-power and SO-phase histograms for all the participants within the healthy controls. Considering the nature of histograms for being distinct, these were also extremely heterogenous and had strong consistency. The two-fold representation provided a robust, high specificity among individuals for improved EEG phenotypes.
The clinical applications for SO-power histograms also suggested new schizophrenia biomarkers leveraging the TF-peak to explore differences between healthy and diseased patients. The results showed a decrease in the spindle activity of data from patients with neurological disorders. Like healthy individuals, schizophrenia patients also showed differences within the subjects and consistency throughout the SO-power and SO-phase histograms for consecutive nights. The unexplored oscillations provide an opportunity for exploration within novel EEG biomarkers.
Final Thoughts
Considering the significance of these transient oscillations, these sleep spindles are quite vital due to their linkage with aging, neurodevelopment, and neurological disorders. The novel characterization in understanding the spindle activity across the continuous sleep oscillation events, the group of researchers has transcended classical pathways. The heterogeneity between the subjects has provided signatures on individual levels and promised new tools for scientific applications such as understanding neuroscience, disease-specific phenotypic traits, and identifying biomarkers for easy characterization and memory consolidation.
Article Source: Reference Paper | Code Availability: TF-peaks
Learn More:
Top Bioinformatics Books ↗
Learn more to get deeper insights into the field of bioinformatics.
Top Free Online Bioinformatics Courses ↗
Freely available courses to learn each and every aspect of bioinformatics.
Latest Bioinformatics Breakthroughs ↗
Stay updated with the latest discoveries in the field of bioinformatics.
Mahi Sharma is a consulting Content Writing Intern at the Centre of Bioinformatics Research and Technology (CBIRT). She is a postgraduate with a Master's in Immunology degree from Amity University, Noida. She has interned at CSIR-Institute of Microbial Technology, working on human and environmental microbiomes. She actively promotes research on microcosmos and science communication through her personal blog.