The significance of comprehending gene expression regulation in tissue development is highlighted by the examination of disease progression through methods such as ChIP-seq and ATAC-seq. On the other hand, the growing volume of data poses computing difficulties, and the dearth of organized tools emphasizes the requirement for effective analysis platforms. Researchers from the Shanghai Institute of Nutrition and Health created the scalable cloud-based Epigenomic Analysis Platform (EAP, https://www.biosino.org/epigenetics) to solve these problems and analyze large-scale ChIP/ATAC-seq data sets effectively. In order to extract biologically significant insights from heterogeneous datasets and automatically produce publication-ready figures and tabular results, EAP uses sophisticated computational algorithms. This allows for thorough epigenomic analysis and data mining in fields like cancer subtyping and therapeutic target discovery.
Introduction
Understanding the regulation of gene expression in tissue development and disease progression is largely dependent on epigenome profiling. Cutting-edge deep sequencing methods like ATAC-seq and ChIP-seq have made it possible to analyze epigenetic variation in disease cohorts and developmental cells, offering important new insights into the processes governing gene expression and the course of disease. However, to explore these massive datasets, researchers need greater computing power as data collection grows in scope. In addition, the large volume of ChIP/ATAC-seq data deposited in NCBI GEO8 and CNCB GSA9 cannot be explored without the use of systematic epigenomic analytic methods. Despite the availability of computational tools and methods, it remains challenging for experimental biologists to deploy and integrate these tools into workable pipelines, particularly in heterogeneous cohort studies where conventional analysis tools are often inadequate.
Understanding EAP (Epigenomic Analysis Platform)
An interactive tool called EAP provides a flexible, adaptable, and scalable way to analyze ChIP/ATAC-seq data. EAP converts enormous data into outcomes that are physiologically meaningful by using sophisticated statistical and computational procedures. Numerous data analytical techniques are available on the platform, such as data preprocessing, supervised differential analysis, enrichment analysis of differential TF motifs, analysis of differential TF activity, unsupervised hypervariable analysis, clustering analysis, and analysis of signature genes scoring. Data mining in a variety of research fields, including cancer subtyping and therapeutic target finding, is made easier by these tools, which are made to model and comprehend epigenetic changes among patient samples or cellular states.
EAP Architecture
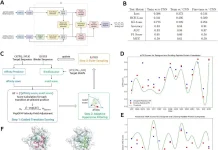
Users must create an account and request storage space on EAP, a cloud-based analytical platform intended for cancer patient cohort studies. A CSV metadata file and input files in FASTQ format are needed for the platform. A md5sum checking process guarantees file integrity, and a specialized data transfer client tool facilitates break-point resumeable transfers. Two analytical modules are available from EAP to convert ChIP/ATAC-seq data into findings that are physiologically significant. It creates an analytical framework for handling, analyzing, and visualizing big datasets by utilizing private cloud computing technology. In order to ensure repeatability and streamline the end-to-end bioinformatics analysis of massive epigenomic datasets, automated analysis pipelines, and tools are created using Docker container technology. Both basic and advanced epigenomic data processing in EAP are supported by the cloud computing architecture.
Application of EAP
- Efficient and comprehensive large-scale ChIP/ATAC-seq data analysis
The Basic Analysis module of EAP is an automated pipeline that handles standard data processing tasks like read alignment, peak calling, read counting, and quality checking. It makes use of cloud computing technology to process ChIP/ATAC-seq datasets (tens or hundreds of profiles from various samples) in an effective manner. The analysis report is provided for additional research, together with quality control plots and summary statistic results. This effective method enhances the quality of NGS data preparation while using fewer computational resources.
EAP is a vast array of analytical tools created to satisfy the demands of personalized ChIP/ATAC-seq data analysis. The Basic Analysis module provides an input read count table that allows users to run both routine and customized studies on their data with ease. Users can set up settings for advanced analyses, such as sample clustering, differential analyses, and p-value/FDR cutoffs, using the easy-to-use interface provided by EAP. The precise set of analytical tools that users need to best fit their study circumstances can be chosen. Differential analysis can identify differences in signals between samples with different labels for ChIP/ATAC-seq datasets that have well-defined sample labels. This can be followed by differential TF activity analysis or differential TF motif enrichment analysis to investigate TFs linked to open chromatin sites or differential genome binding. For datasets with no predefined sample labels or highly sophisticated sample labels, hypervariable analysis can identify hypervariable ChIP/ATAC-seq signals across the samples, which can be used for clustering analysis to dissect the underlying heterogeneity structure among the samples.
- Provides interactive data set browser
EAP has successfully examined ChIP/ATAC-seq data from a number of epigenomic studies on cancer, including the LUAD cohort, NSCLC cohort, thyroid cancer cohort, TCGA pan-cancer cohort, and organoids produced from pancreatic cancer patients. Users can explore the involvement of transcriptional regulators in oncogenesis by accessing the data sets through the Data Set Browser in EAP, which offers an interactive interface for simple visualization of TF activity scores in each data set.
Conclusion
Large-scale investigations of epigenomic datasets can be supported by the cloud-based data analysis platform EAP. It has proven to be highly effective in analyzing the epigenetic heterogeneity of cancer, identifying transcription factors linked with specific cancer subtypes, and identifying important transcriptional regulators of various cancer clinical subtypes. With the help of interactive data analysis provided by EAP, users can improve the identification of biologically meaningful results by iteratively changing analytical settings. It has been effectively used to examine ChIP/ATAC-seq data from multiple epigenomic studies on cancer, demonstrating increased regulation activity in metastatic tissues as well as progressive subtypes. The RUNX family of transcription factors showed increased regulation activity in metastatic tissues as well as progressive subtypes, indicating that they could be useful targets for cancer treatment. EAP is expected to be used for more large-scale cancer epigenomic data analyses, leading to more interesting and meaningful biological or clinical discoveries and broadening our understanding of cancer carcinogenesis and progression-related epigenetic machinery.
Article Source: Reference Paper | The software is freely available as open source, with its entire code accessible on GitHub.
Important Note: bioRiv releases preprints that have not yet undergone peer review. As a result, it is important to note that these papers should not be considered conclusive evidence, nor should they be used to direct clinical practice or influence health-related behavior. It is also important to understand that the information presented in these papers is not yet considered established or confirmed.
Follow Us!
Learn More:
Deotima is a consulting scientific content writing intern at CBIRT. Currently she's pursuing Master's in Bioinformatics at Maulana Abul Kalam Azad University of Technology. As an emerging scientific writer, she is eager to apply her expertise in making intricate scientific concepts comprehensible to individuals from diverse backgrounds. Deotima harbors a particular passion for Structural Bioinformatics and Molecular Dynamics.