Conventionally, cancer diagnostic procedures have been known for their invasive nature and enormous cost. However, researchers from KAIST and Korea University have developed a novel diagnostic approach that combines AI and innovative technology for an affordable, accessible, and non-invasive diagnosis of multiple types of cancer at an early stage under high resolution. In the study, small vesicles released by cancer cells were analyzed using Exosome-SERS and AI to achieve promising outcomes in the early detection of various types of cancers with a high degree of accuracy. This breakthrough study could significantly impact cancer diagnosis and monitoring, transforming current practices and offering optimism for cancer patients.
Utilizing exosomes for cancer diagnosis
Cancer is a devastating disease group responsible for many deaths worldwide. Early diagnosis, which can also enhance therapy choices for many patients, can considerably increase the likelihood of survival. Sadly, no one technique can now reliably and efficiently identify a variety of early-stage tumors in a non-invasive manner. For the early detection of cancer, current biomarkers such as prostate-specific antigen (PSA) and carcinoembryonic antigen (CEA) are neither sensitive nor specific enough. Moreover, other cancer types have no screening method for their detection.
The study published in Nature Communications proposes a unique approach to overcome this challenge by using exosomes, AI, and surface-enhanced Raman spectroscopy (SERS). Exosomes are small vesicles excreted by living cells, which carry information about their origin and condition. These can be found in a variety of body fluids, such as blood, urine, cerebrospinal fluid, and saliva. These vesicles are essential for a number of physiological processes, including those that influence the behavior of cancer cells, control the body’s immunological response, and encourage the development of new blood vesicles. Since exosomes are also indicative of the molecular characteristics of their parent cells – which could even be oncogenic/cancer cells – they can be used as biomarkers for cancer diagnosis and prognosis.
Yet, analyzing exosomes represents a significant challenge. Detecting and characterizing exosomes is not a simple process due to their small size of roughly 0.1µm in diameter and their heterogeneity. The limitations of conventional techniques like ELISA and PCR are due to the requirement of certain antibodies or primers that target particular components in exosomes. This limits the sensitivity and feasibility of these techniques. Secondarily, these methods consume time, are labour-intensive, and are expensive.
SERS technology and AI
Combining SERS technology and AI can help overcome the challenge of using exosomes for cancer diagnosis and monitoring. SERS, a molecular analysis technique, utilizes metal nanoparticles as substrates to amplify the Raman scattering signals of molecules. Raman scattering happens when light interacts with a substance, causing the scattered light to change frequency. The specific changes in frequency depending on the molecular structure and composition of the substance. Essentially, analyzing the scattered light that comes back after shining light on the sample allows the system to learn about the substance’s chemical composition. By creating hotspots of electromagnetic fields on metal nanoparticles’ surfaces, SERS acts as an incredibly sensitive and powerful tool for biochemical identification and analysis of low-concentration heterogeneous molecules, like exosomes, with high resolution. Essentially, because different types of exosomes have unique spectral fingerprints, they can be differentiated using SERS.
Artificial intelligence, by using machine learning algorithms, can help analyze the complex and large-scale data produced by SERS and can, subsequently, make predictions or classifications based on patterns or features. Implementation of AI can also minimize human error and bias, thereby augmenting the accuracy and efficiency of SERS-based diagnosis.
Effectiveness of the single test diagnostic tool
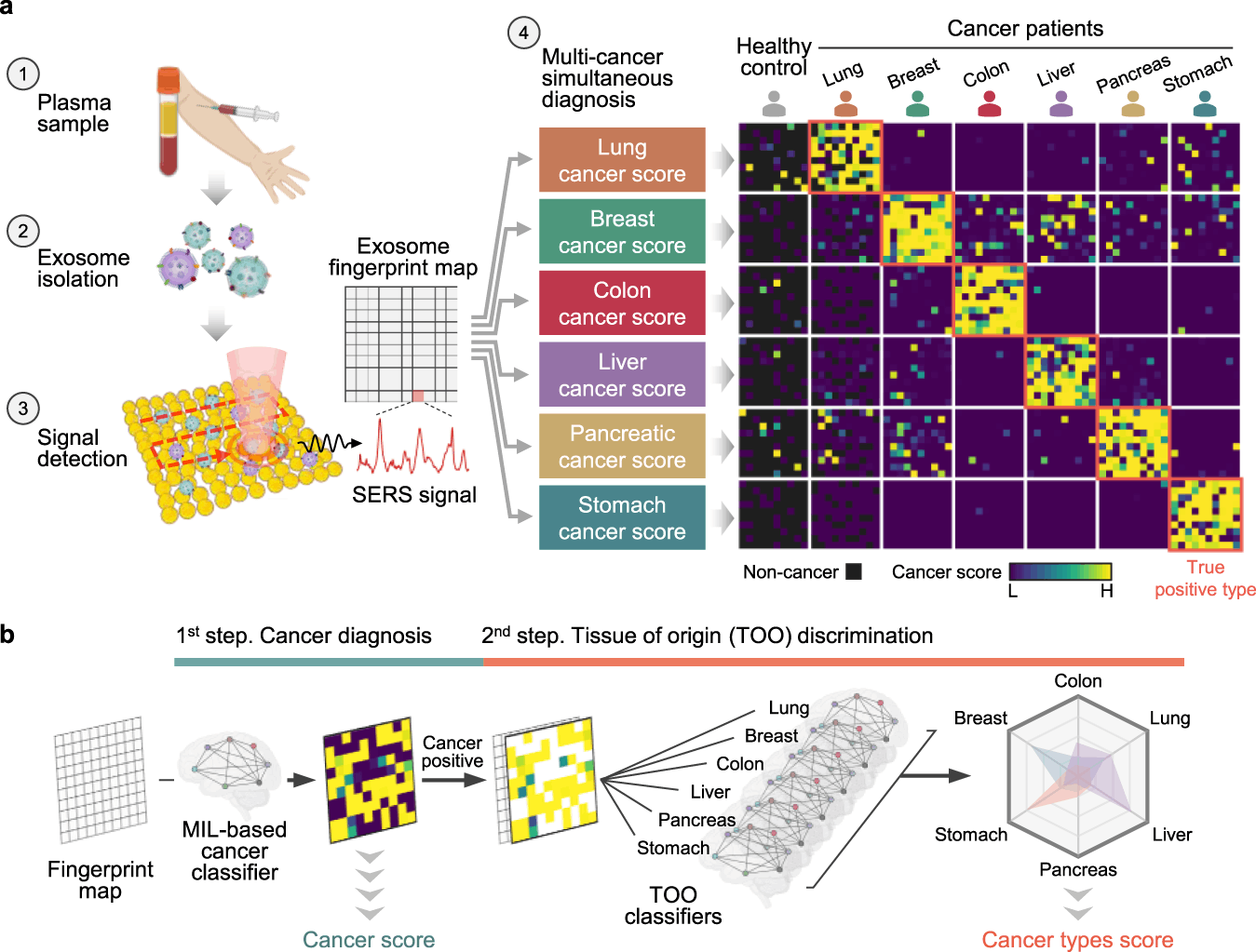
The researchers have introduced a diagnostic tool that can use AI to analyze the SERS profiles of exosomes from plasma samples to simultaneously identify six types of early-stage cancers – lung, breast, colon, liver, pancreas, and stomach. The diagnostic procedure of the system comprises three major steps; firstly, it conducts isolation and enrichment of exosomes from plasma via gel filtration chromatography. Secondly, the exosomes are labeled with gold nanoparticles, and their SERS spectra are measured. Lastly, exosomes are grouped either into cancer or healthy and are then identified based on their tissue of origin using machine learning models.
The single test-based diagnostic tool was assessed on 520 plasma samples from healthy individuals and patients pre-diagnosed with different stages of cancer. The findings were remarkable because an area under the curve (AUC) value of 0.970 was obtained for cancer detection. The tool is highly accurate in distinguishing between cancer and normal samples. In addition, the researchers achieved a mean AUC of 0.945 for tumor organ type classification, highlighting the fact that this system can precisely detect the specific tissue of origin of cancer exosomes. The final integrated decision model demonstrated a sensitivity of 90.2% and a specificity of 94.4% for cancer detection while also successfully predicting the tumor organ type for 72% of positive patients. These findings are crucial and highlight the diagnostic system’s great potential for improving cancer diagnosis, especially at an early stage.
The study highlights that the system has several distinct advantages compared to other existing methods for cancer diagnosis. First, the system’s non-specific analysis of Raman signatures means it does not rely on pre-characterized biomarkers/antibodies, which can vary depending on the cancer type or stage. Additionally, its ability to identify six types of early-stage cancers with a single test potentially reduces the cost and time required for screening and diagnosis. Lastly, as discussed previously, the system uses AI to learn from existing data and train itself to improve over time.
Despite the various advantages discussed, some limitations and challenges of the study prevail. For instance, the results must be validated on more extensive and diverse cohorts of patients with other diseases or conditions that could affect exosome properties and SERS spectra. The system must also be improvised and optimized for clinical settings. This involves improving the sample preparation process, minimizing the background noise and interference from other matter, and validating the reproducibility and robustness of the findings and models.
Conclusion
To conclude, the study presents a promising tool for the early detection of multiple cancer types using exosomes, SERS, and AI. The system could positively impact cancer patients’ outcomes and quality of life by allowing timely and accurate diagnosis. However, more research is required to validate the method’s viability in real-world scenarios.
Article Source: Reference Paper
Learn More:
Diyan Jain is a second-year undergraduate majoring in Biotechnology at Imperial College, London, and currently interning as a scientific content writer at CBIRT. His passion for writing and science has led him to pursue this opportunity to communicate cutting-edge research and discoveries engagingly to a broader public. Diyan is also working on a personal research project to evaluate the potential for genome sequencing studies and GWAS to identify disease likelihood and determine personalized treatments. With his fascination for bioinformatics and science communication, he is committed to delivering high-quality content a CBIRT.