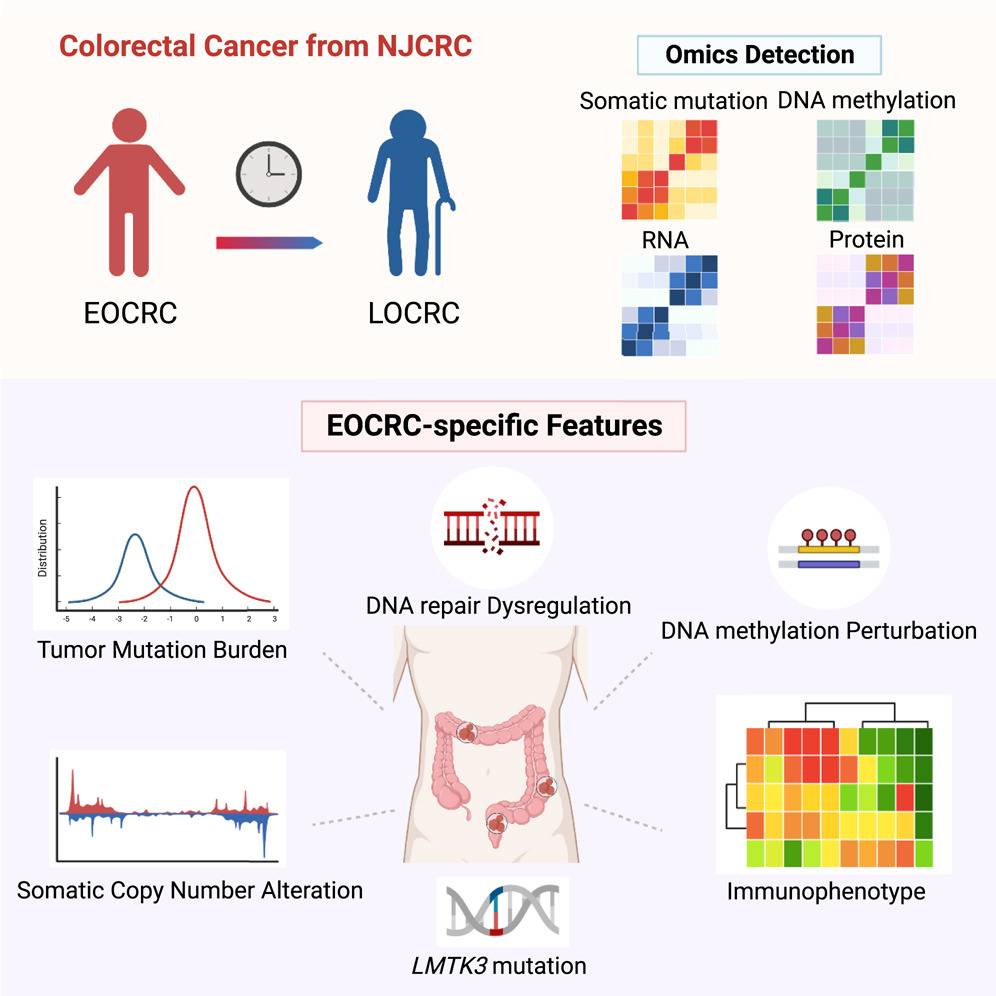
There is a growing number of incidences of colorectal cancer in people under the age of 50 (EOCRC), but the intrinsic molecular mechanisms underpinning it are not very well understood. A recent study was conducted on 79 CRC patients of Chinese ancestry to understand the differences in the omics of EOCRC compared to general CRC. The study presents incredible findings and shows that EOCRC has many tumor mutations, increased DNA repair features, and upregulated immune infiltration. The authors also discovered a potential biomarker, LMTK3, that may functionally drive EOCRC development and immunotherapy. The study also provides crucial information for precision oncology in treating EOCRC.
Colorectal Cancer and the Need for a Comprehensive Understanding of its Etiology
Colorectal cancer is a type of cancer that affects the colon and rectum. It is the third most common cancer type and the second most deadly, and therefore significantly impacts global health. While developed countries have seen a decrease in colorectal cancer cases due to advancements in cancer screening/diagnosis technology, there has been a rise in early-onset colorectal cancer (EOCRC), particularly in China, making it an issue that must be addressed.
Various established risk factors for late-onset colorectal cancer (LOCRC) comprise consumption of a Western-style diet, obesity, physical inactivity, and molecular alterations in driver genes of APC, TP53, and KRAS. However, patients with EOCRC possess distinct features compared to those with LOCRC, including male predominance, less environmental exposure accumulation, a history of inflammatory bowel disease or a family history of colorectal cancer, and tumors on the left side with molecular and genetic heterogeneity.
Furthermore, research has demonstrated a higher tumor mutation burden (TMB) in African American patients with EOCRC than in White patients. Also, mutational frequencies in driver genes differed between diverse populations of patients with EOCRC and LOCRC. These studies suggest a differential mutational landscape across various ages of onset and diverse ancestries. Moreover, previous studies have also described multi-omics features of EOCRC to understand the molecular basis of the disease. However, these studies conducted so far have only focused on patients who are not of Asian descent, implying the need for further research to fully understand the etiological factors that contribute to the development of colorectal cancer at a specific age of onset.
A Multi-Omics Approach to Studying EOCRC Patients of Chinese Ancestry
A paper published on Cell Reports Medicine recently reports the genomic, epigenomic, transcriptomic, and proteomic profiles of EOCRC tumors based on a Chinese colorectal cancer cohort. The authors of the paper aimed to identify the distinct multi-omics architecture of EOCRC tumors compared to LOCRC tumors to better understand age-of-onset disparities in colorectal cancer etiology and provide valuable insights into developing precision therapies.
The study included 79 patients with colorectal cancer, with 30 patients having EOCRC and 49 having LOCRC. Both tumor and healthy tissues were collected for multi-omics profiling. The chronological age of diagnosis of the patient positively correlated with their DNA methylation age in tissues. Moreover, the transcriptomic profiles between EOCRC and LOCRC patients were compared, and the pathway enrichment analysis demonstrated that the top 10 KEGG pathways in either tumors or NATs included pathways related to aging-associated diseases. These initial findings were crucial and illustrated the ability of a multi-omics approach to identify EOCRC-specific molecular features.
EOCRC-Specific Molecular Features Determined by Multi-Omics Data and Development for New Therapies for EOCRC
Transcriptomic and proteomic profile analyses showed that fatty acid degradation (FAD), cell adhesion molecules (CAMs), and DNA repair pathways were involved in EOCRC development. ADH1B, a crucial enzyme in alcohol metabolism, was downregulated, and CADM3, an essential protein in the CAM pathways, was reduced in EOCRC tumors. A significant increase in signature 3, associated with DNA repair, was seen in EOCRC tumors, indicating a crucial interplay between the immune-inflammation response and DDR in the young tumor microenvironment. These identified molecular phenotypes greatly enrich our understanding of EOCRC’s etiology.
Distinct omics profiles offered valuable insights into the development of new therapeutic methods for EOCRC. Given that EOCRC tumors have substantially high TMB and remarkably unique pathway enrichment in mismatch repair, immunotherapy would be the preferred therapeutic approach. The study identified potential drug targets by profiling the gene expression spectrum for platinum drug resistance, CT antigens, pan-essential genes, and an immune panel. It was observed that most genes demonstrated inconsistent transcriptomic and proteomic expression patterns in the comparison between age-of-onset subgroups, which potentially attributed to tighter post-transcriptional regulation in young tumors. The authors, thereby, took a precision approach to drug development and therapeutic treatment/management for EOCRC.
Furthermore, LMTK3, a protein kinase modulating ERα, varied in mutation frequency across age-of-onset and ancestry, and this is likely to have triggered molecular alterations in EOCRC tumorigenesis. EOCRC mutant LMTK3 upregulated gene expression in defined oncogenic pathways, consistent with the molecular phenotypic features of EOCRC. Two DNA repair genes – SHPRH and POLE – were mutated with LMTK3, corresponding to the increased signature 3, as discussed. LMTK3 is, hence, a druggable target for colorectal cancer therapy, independent of its mRNA expression.
Conclusions and Limitations
To conclude, the study performed a molecular characterization of colorectal cancer through multi-omics data analysis approaches, including genomic, epigenomic, transcriptomic, and proteomics, underpinning the molecular complexity and heterogeneity based on differences in the age of onset and ancestry. The findings are crucial and can be used to mitigate disparities among patients with this type of cancer by examining precision oncology in drug development and therapeutic modalities.
However, there are a few limitations of the study that need to be addressed through future work:
- Only a limited number of samples were sequenced; therefore, the authors could only determine some obvious molecular alterations but significant ones.
- The authors did not explore PTMs like glycomics or phosphoproteomics in their multi-omics data. Addressing these two limitations could provide further insights into the molecular heterogeneity of EOCRC.
- The authors did not conduct survival analysis or experimental validation of the LMTK3 mutation because of the short-term follow-up in the CRC cohort.
Hence, larger sample sizes, deeper profiling, and functional experiments could greatly augment our knowledge of CRC etiology and pathophysiology.
Article Source: Reference Paper
Learn More:
Diyan Jain is a second-year undergraduate majoring in Biotechnology at Imperial College, London, and currently interning as a scientific content writer at CBIRT. His passion for writing and science has led him to pursue this opportunity to communicate cutting-edge research and discoveries engagingly to a broader public. Diyan is also working on a personal research project to evaluate the potential for genome sequencing studies and GWAS to identify disease likelihood and determine personalized treatments. With his fascination for bioinformatics and science communication, he is committed to delivering high-quality content a CBIRT.