Scientists from Inserm, Université Sorbonne Paris Nord, and Université Paris-Cité at the IAME Laboratory, with their collaborators at Institut Pasteur and the Paris Public Hospitals Group (AP-HP), develop an in silico model in combination with in vivo phage therapy data to highlight vital parameters for increasing the efficiency of pneumonia treatment.
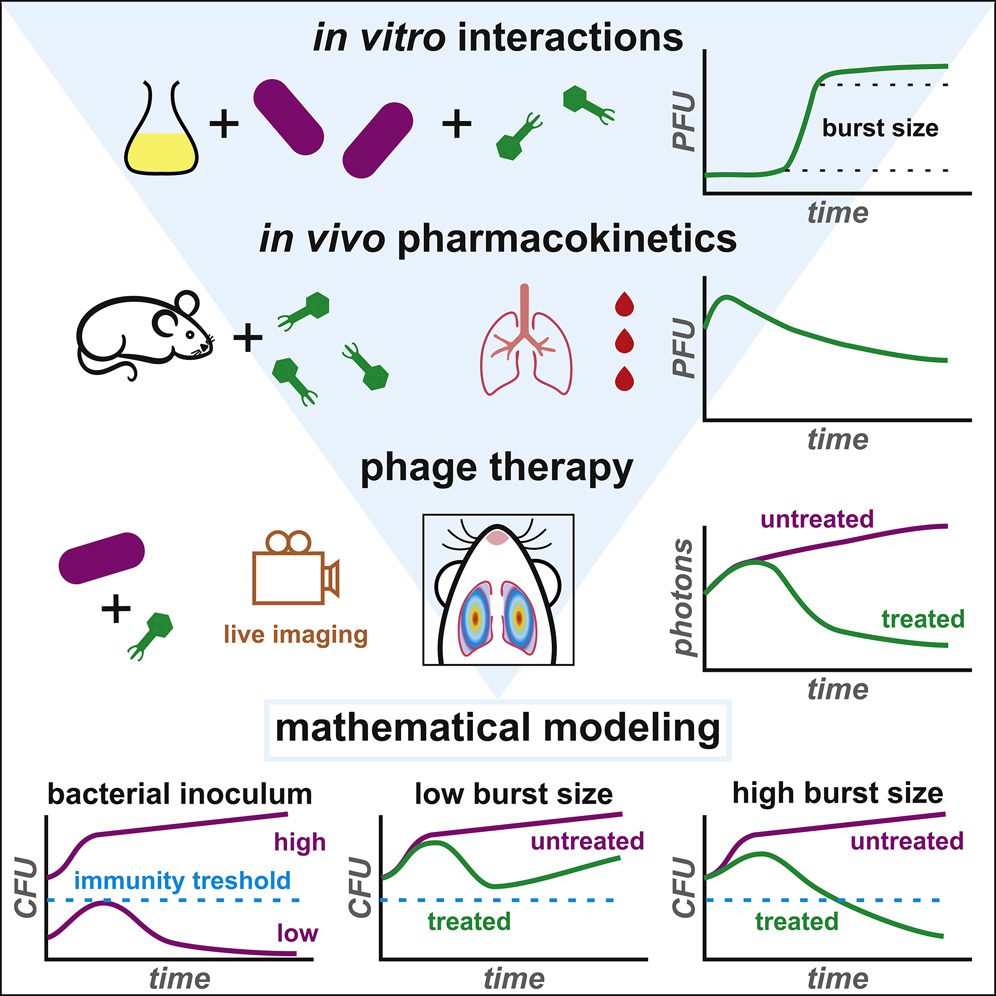
Image Source: Combination of in vivo phage therapy data with in silico model highlights key parameters for pneumonia treatment efficacy
The clinical (re)development of bacteriophage therapy for the treatment of antibiotic-resistant diseases incurs the difficulty of understanding the dynamics of the interaction between phage and bacteria in the in vivo context.
Described in the paper is a general strategy coupling in vitro and in vivo experimentations with a mathematical model for the characterization of the interplay among phage and bacteria during pneumonia prompted by a pathogenic strain of Escherichia coli.
The model permits the assessment of a few vital parameters for phage therapeutic efficacy. Specifically, it measures the effect of dose and route of phage administration as well as the synergism of phage and the innate immune response on bacterial clearance.
Simulations predicted a restricted effect of the intrinsic phage characteristics in concurrence with the ongoing semi-empirical choices of phages for compassionate treatments. Model-based approaches would foster the deployment of future phage-treatment clinical preliminaries.
Successes and Failures
The articulated slowdown in the discovery of new antibiotics for the treatment of bacterial diseases brought about by multidrug-resistant (MDR) microorganisms has renewed interest in bacteriophages which are viruses that infect bacteria.
Pre-clinical experimentations and numerous clinical case reports have exhibited the capability of phages to effectively treat infections brought about by MDR bacteria.
These successes are regardless balanced by unreported failures as well as disappointing clinical examinations, showing that interpretation from experimental outcomes to clinical preliminaries stays challenging.
Significant Achievement of Widespread Usage of Phages in Treatment
A considerable achievement for the far and wide use in human populaces is the determination of the ideal dosage, administration route, and treatment duration.
This is an especially intricate undertaking for phages, as standard evaluations of clinical pharmacology used for the determination of the cycles of administration, distribution, metabolism, and excretion (ADME) of drugs are not adapted to phages.
To be sure, phages hugely replicate on their target bacteria, with kinetic parameters administering this viral amplification (receptor recognition, genome amplification, particle assembly, and bacterial lysis) being well defined for every phage-bacterium pair.
Besides, phages are generally present in human-related microbiotas, and their rate and route of elimination don’t follow common metabolic pathways of xenobiotic products through the kidney or liver.
A New Study
In this study, the scientists coupled in vitro and in vivo experimentations with mathematical modeling to move toward the critical parameters of phage-bacteria-host interactions and guide treatment strategies.
This approach has been fruitful in drug improvement for different diseases such as HIV, hepatitis C virus, and acute diseases like Ebola, Zika, and Influenza, which are viral diseases.
Dissimilar to past models that focussed on specific parts of phage-bacteria, phage-host immune system, or bacteria-host immune framework interactions, in this study, the scientists had an objective to design a synthetic, semi-mechanistic model that could be utilized to comprehend the tripartite phage-bacteria-host interactions in vivo.
The scientists designed explicit experimentations to estimate the vital parameters of phage-bacteria interactions by utilizing a formerly characterized experimental model.
The virulent phage 536_P1, a Myoviridae belonging to the Phapecoctavirus class, rapidly (20 min) lyses the hypervirulent Escherichia coli strain 536 in vitro.
Besides, a solitary dose of phage 536_P1 administered intranasally to mice 2 hours post-infection by a lethal dosage of strain 536 saved 100 percent of the animals, while untreated animals died within a 72-hour bracket, showing the efficiency of this phage in treating an acute pulmonary infection.
In the study, dynamic and direct microbiological information was gathered from groups of animals that were either bacterial uninfected and phage treated, infected and untreated, or infected and treated, utilizing different inoculum dosages and routes of administration.
This information drove the researchers to foster a unique mathematical model restating the tripartite interactions over time. This model identified vital parameters influencing bacterial kinetics in vivo.
It was then taken advantage of to test situations, including varieties of phage intrinsic properties. The utilization of such a model would rationalize the phage choice, the dosage, and the route of administration to streamline the efficiency of phage treatment.
Inspiration for the Study
The experimentation study was inspired by the work performed during the improvement of new drugs, where longitudinal pharmacokinetic and pharmacodynamic information were gathered to figure out the exposure-response relationship and optimize the dosage routine.
It is critical to recognize that bioluminescence is, in any case, an intermediary of bacterial load compared with CFU counts from lung samples, as reflected by the estimation error related to bioluminescent information.
Nonetheless, to a great extent, this estimation is remunerated by longitudinal recordings that give a unique insight into the determinants of bacterial kinetics for every animal.
The heterogeneity of the underlying bacterial loads noticed 2 hours post-infection gave an enormous scope of circumstances reflecting clinical situations.
The assortment of individual profiles was captured by utilizing non-linear mixed effect models to estimate phage-bacterial interaction. This statistical methodology gets strength from the between-subjects variability to increase the accuracy of parameter assessment and coax out the impacts of bacteria parameters, dosages, and routes of administration.
An immediate result of the above conditions was the exact measurement of the threshold value (6.94 log10 CFUeq/g), above which the immune response was overpowered without a trace of phage therapy.
To guarantee that the model restates the kinetics of phage-bacteria interactions during treatment, it was essential to use, however much as could reasonably be expected, parameters deduced from animal experimentations rather than in vitro conditions.
For example, the bacterial growth rate, infectivity rate, and lysis rate were undeniably reasoned from experimental information. In any case, a few parameters ought to be all the more profoundly explored to work on the fidelity of the model.
In this work, the scientists decided to represent the immune response in a synthetic manner that envelops mechanical factors, like mucins and surfactant proteins, as well as different cell actors of the innate immune system.
In any case, the immune response would have been all the more mechanistically characterized if a few markers (cytokines, immune cells) had been monitored. This model likewise considers the lung compartment as homogeneous, which overlooks the complex structure of the organ that will probably influence phage-bacteria interactions.
The Endpoint
To summarize, the model created during this work could be utilized to predict the efficiency of basically any phage for which a minimum set of in vitro and in vivo information should be acquired, which will impressively bring down the number of experimentations expected to validate such a phage during pre-clinical development.
Past phage treatment, the model could likewise be carried out to test combined anti-infectious treatments like the association of phages with antibiotic agents.
Article Source: Delattre, R., Seurat, J., Haddad, F., Nguyen, T.-T., Gaborieau, B., Kane, R., … Debarbieux, L. (2022). Combination of in vivo phage therapy data with in silico model highlights key parameters for treatment efficacy. Cell Reports, 39(7). https://doi.org/10.1016/j.celrep.2022.110825
Learn More:
Top Bioinformatics Books ↗
Learn more to get deeper insights into the field of bioinformatics.
Top Free Online Bioinformatics Courses ↗
Freely available courses to learn each and every aspect of bioinformatics.
Latest Bioinformatics Breakthroughs ↗
Stay updated with the latest discoveries in the field of bioinformatics.
Tanveen Kaur is a consulting intern at CBIRT, currently, she's pursuing post-graduation in Biotechnology from Shoolini University, Himachal Pradesh. Her interests primarily lay in researching the new advancements in the world of biotechnology and bioinformatics, having a dream of being one of the best researchers.











