Researchers from Pennsylvania State University have put forth a novel machine-learning approach to help predict Autism Spectrum Disorder in children at a very young age, between 18 to 30 months. Such early detection can vastly help both the patients and the clinicians to improve treatment outcomes.
The authors define Autism Spectrum Disorder (ASD) as a developmental disability. ASD arises due to differences in the brain, which indicates a neurological origin. There is no single cause of autism; given the complexity of the disorder, both nature (genetics) and nurture (environmental factors) seem to play a role. Genetically, one of the significant types of autism disorder called Rett’s syndrome is caused by mutations on the X chromosome. Environmentally, factors such as microbial infections, air pollution, and complications during pregnancy also seem to play a role in triggering ASD.
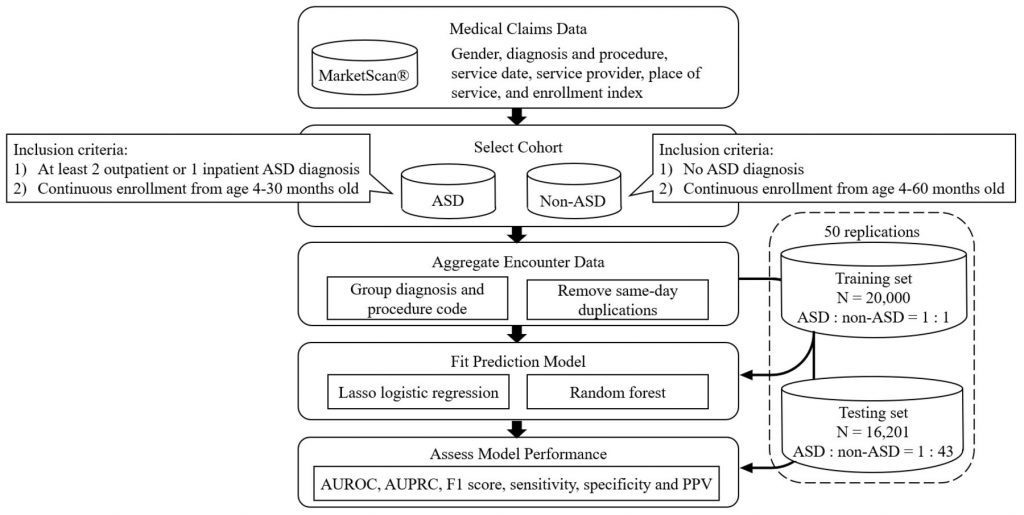
Image Source:http://dx.doi.org/10.1136/bmjhci-2022-100544
People with ASD also face severe challenges in social interaction, poor communication skills, and an inability to identify non-verbal cues. They also show signs of limited, severe, repetitive behavioral patterns. According to the Centre for Disease Control’s (CDC) Autism and Developmental Disabilities Monitoring (ADDM) Network, an eighteen-year-long survey identified about 1 in 44 children around the ages of 8 with ASD in 2018. The children face long-term struggles maneuvering through daily life, education, and employment, but early intervention and diagnosis can significantly help improve the symptoms.
According to CDC, children about two years are when a reliable diagnosis of ASD can be made. To enhance early ASD diagnoses, the American Academy of Pediatrics (AAP) has recommended using the Modified Checklist for Autism in Toddlers (M-CHAT), a 20-question test to evaluate ASD in toddlers aged 16 to 30 months (with 18 to 24 months being the perfect window time). A growing body of evidence suggests M-CHAT alone might be a poor screening tool with very low sensitivity, as ASD-specific behavioral patterns often underpin medical issues such as gastrointestinal problems, viral infections, and feeding issues. Thus, relying only on M-CHAT can result in misdiagnosis of the child. Therefore to avoid misidentification, children are not officially diagnosed until they are four or five years old, missing the potential screening time.
To fill this gap, a team of interdisciplinary researchers led by Qiushi Chen, assistant professor at the Penn state college of engineering, have published their novel finding in the BMJ Health and Care Informatics. The study has been funded by the National Institutes of Health, the Penn State Social Science Research Institute, and the Penn State College of Engineering to help develop a computational model that utilizes the Electronic Healthcare Record (EHR) of previous clinical visits to help identify ASD risk in toddlers.
Findings
Professor Chen states that the key idea is to improve how the researchers utilize the resources, with a team of clinical and computational experts to develop a tool that physicians can use without any specialized knowledge to aid in diagnosing children as early as possible.
The authors used the IBM MarketScan Health Claims database containing health insurance claim data of 273 million unique individuals across the USA from 2005 to 2016. But the study cohort consists only of 12,743 young children with ASD and 25,833 random samples of young children without ASD. The study implied risk-predicting machine learning models such as logistic regression (LR) and random forest (RF), out of which RF showed increased performance than LR. The model showed an overall AUROC (Area Under the Receiver Operating Characteristics) above 0.75 when predicting ASD at 24 months.
The results have shown that the model’s prediction performance significantly increased with age-with an AUROC of 0.717 at the age of 18 months to an AUROC of 0.832 at the age of 30 months. Clinically interpretable factors are considered to be key predictors in the study. Predictors including sex (male, since they are more frequently diagnosed with ASD), developmental delays, clinical visits for gastrointestinal disorders, respiratory system infections, and otitis media (middle ear infection and antibiotic usage is linked to autism) are strongly correlated to ASD outcomes. The study has also shown that separating in-patient from out-patient claims as a predictor for ASD further increased the model’s accuracy with a specificity of 96.4% at a sensitivity of 40%. This outperformed the already available M-CHAT/F, with a reported sensitivity of 38.8% and specificity of 94.9%.
Conclusion
Chen et al. have demonstrated the proposed model’s predictability to assess Autism Spectrum Disorder in toddlers using the EHR system. The model was also shown to be clinically interpretable with systemic identification of key predictors, which is in line with Autism Spectrum Disorder symptoms and risk factors reported in the literature. Since the ML model utilizes the medical history database only, it is feasible to integrate the model with the database, thereby making it an automatic screening tool. Detailed clinical descriptions in the database can enable the tool to be used more than a mere screening tool to increase accuracy; instead can be used to direct screening decisions.
The model does suffer from a critical limitation where the absence of any ASD risk factors in a child’s medical record does not imply that the child cannot or will not develop ASD. Despite this, the model has shown robust outcomes and accuracy. As the early diagnosis of ASD is crucial to improve the lives of the patients, the ability of the proposed study to detect ASD risk as young as 24 months, as opposed to four or five years, can greatly impact treatment outcomes. A similar approach can also be taken in the near future to assess and predict several other disease implications in the population.
Article Source: Reference Paper | Reference Article
Learn More:
Top Bioinformatics Books ↗
Learn more to get deeper insights into the field of bioinformatics.
Top Free Online Bioinformatics Courses ↗
Freely available courses to learn each and every aspect of bioinformatics.
Latest Bioinformatics Breakthroughs ↗
Stay updated with the latest discoveries in the field of bioinformatics.
Shwetha is a consulting scientific content writing intern at CBIRT. She has completed her Master’s in biotechnology at the Indian Institute of Technology, Hyderabad, with nearly two years of research experience in cellular biology and cell signaling. She is passionate about science communication, cancer biology, and everything that strikes her curiosity!







