With the potential to have significant impacts on various fields, like therapeutics and agritech, protein engineering has recently been a subject that has incited much intrigue within the scientific community. However, despite this, many factors that govern the synthesis and regulation of proteins aren’t well-known. The intricate relationships that influence the various interactions proteins have with each other and their environment at large are so numerous that they form a seemingly intractable web. However, new research utilizing deep learning provides a potential new method of using amino acid sequences to predict the abundance of proteins.
Despite decades of research into the area, our understanding of amino acid sequences’ role in proteome abundance is still relatively scarce. The regulation of protein synthesis is complex and often involves the confluence of multiple factors. Protein synthesis and degradation occur in tandem to adjust protein levels, and their interplay has important implications for genetic and protein engineering. However, there are indications that the majority of the regulation occurs primarily at the degradation stage and is partially encoded inside of the protein’s amino acid sequence.
On a larger scale, proteins that play key roles in cellular functions or serve as essential nodes in interaction networks frequently have higher abundances. Due to their potentially significant effects on cellular fitness, these extremely abundant proteins are subject to strict evolutionary restrictions and evolve more slowly than other proteins. Surprisingly, steady-state protein abundance persists across numerous evolutionary lineages, from microbes to mammals. According to theoretical models, decreasing fitness caused by an increase in protein abundance slows evolution, with the least stable proteins being able to adapt faster. However, proteins can evolve more quickly by acquiring mutations that improve their stability and folding. Additionally, experimental data suggests that the mutational robustness afforded by additional protein stability enhances a protein’s ability to evolve, increasing evolvability. This allows the protein to be able to accept a larger range of advantageous mutations even as it folds to a native structure.
Highly expressed proteins are frequently more thermostable, which is commonly explained by the so-called misfolding avoidance hypothesis. Considering that stable proteins are evolved to endure translational mistakes. In contrast, numerous empirical studies showed no significant indication that protein stability and abundance correlate. Additionally, it is known that compared to the metabolic cost of synthesis, the cost (in terms of misfolded protein) of translation-induced misfolding is minimal, hence demonstrating that MAH cannot account for the delayed evolution of highly abundant proteins.
The Use of A Neural Network to Reveal Biological Relationships
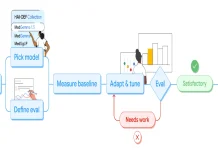
A new study attempts to reconcile these differences using a novel deep learning approach: a deep neural network transformer (also known as BERT) was developed and trained using data extracted from 21 previously conducted studies of the proteome. Utilizing this network, over 50% of protein copy number variations could be predicted in yeast cells, using information solely from the amino acid sequence of those proteins. The protein abundances of more than 5000 proteins were estimated. The neural network has a self-attention process, which, when studied, revealed that the network had been able to identify specific physicochemical properties that were encoded within the amino acid sequences and which correlated to the conformational stability of the protein. Since the model was trained from scratch, the interpretability of the results was significantly increased.
Mutation Guided by an Embedded Manifold (MGEM) was introduced in order to investigate the sequence, and it was found that mutations that enhanced abundance had significant effects on the protein’s hydrophobicity and polarity. Molecular dynamics simulations were also performed to further lend credence to the theory that stability and abundance are connected. Proteomics experiments were conducted on yeast, revealing that mutated proteins were more abundant than wild-type variants. Crucially, these mutants were found to have lower synthesis costs than the native proteins, showing that it is also advantageous to cellular fitness. While this doesn’t necessarily indicate a direct link between cost and abundance, it shows that the model was able to pick up on such information through only amino acid sequences.
Prior studies showed that physicochemical properties like polarity affect a protein’s stability and translational speed. Links to backbone conformation, as well as a marked preference for alpha helix, suggest that there may be a link to the secondary structure of the protein as well. It was also found through mutants created by MGEM that the N-terminus is of great importance when predicting protein abundance. There weren’t many changes in the region, which indicates that it is optimized for efficiency of expression. While it is true that mutations can cause significant structural changes that may destabilize proteins, it should be noted that changes in backbone conformation alone do not necessarily indicate protein stability. To gain a more comprehensive understanding of this phenomenon, an investigation was conducted into intermolecular interactions, specifically focusing on the number of contacts between neighboring amino acids. The findings suggest that proteins that are prone to denaturation tend to expose their hydrophobic core, leading to a loss of hydrophobic interactions and an increase in solvent accessibility. To further explore the effects of substitutions on hydrophobic cores, the Solvent Accessible Surface Area (SASA) was computed for all proteins under consideration. The results indicated a significant decrease in SASA for abundance-increasing mutants compared to wild types, thus supporting the hypothesis.
Conclusion
Analysis utilizing deep neural networks shows great promise in uncovering new knowledge of proteins. While the study’s primary aim was to demonstrate relationships between protein abundance and its amino acid sequence, important revelations were produced regarding the influence of factors such as conformational stability and metabolic cost. Remarkably, both methods optimized sequences for metabolic cost without needing to be conditioned explicitly, demonstrating that such knowledge can be gained purely through analysis of sequence data, showing the significant potential of deep learning in protein engineering.
Article source: Reference Paper
Important Note: bioRiv releases preprints that have not yet undergone peer review. As a result, it is important to note that these papers should not be considered conclusive evidence, nor should they be used to direct clinical practice or influence health-related behavior. It is also important to understand that the information presented in these papers is not yet considered established or confirmed.
Learn More:
Sonal Keni is a consulting scientific writing intern at CBIRT. She is pursuing a BTech in Biotechnology from the Manipal Institute of Technology. Her academic journey has been driven by a profound fascination for the intricate world of biology, and she is particularly drawn to computational biology and oncology. She also enjoys reading and painting in her free time.