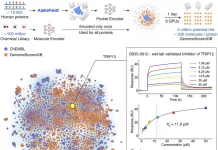
The factors at play in the binding of proteins are intricate and hard to untangle – despite great advancements in recent years in humanity’s understanding of the proteome, conventional methods for determining binding sites and uncovering the mechanisms that influence small molecule-protein binding remain difficult, resource-intensive, and expensive. Computational biology may provide a more efficient and effective solution.
Determining the precise sites at which proteins interact with small molecules helps reveal valuable information regarding the mechanisms that underlie the use of bioactive compounds and also helps in optimizing the selectivity and potency of various compounds. Many kinds of structural techniques like nuclear magnetic resonance and X-ray crystallography have been used for this purpose, but they are limited as they only work with purified and well-behaved targets. Furthermore, these techniques aren’t able to properly capture the interactions of proteins with small molecules in dynamic, practical environments where unpredictable influences like posttranslational modifications, localization, and protein-protein interactions, among others, can change the structure and function of the protein, along with their ability to bind to the molecules above. These methods also don’t adequately depict the interactions of these molecules across the wider proteome and don’t facilitate discoveries of novel binding sites on these proteins.
In order to bridge these gaps, a variety of methods have been created specifically for mapping protein-small molecule interactions within native systems. Photoaffinity labeling is often utilized in order to preserve these interactions and involves a photoreactive group being attached to a given compound, which, when exposed to ultraviolet light, results in the creation of intermediates that are highly reactive and quickly bond with adjacent proteins. Though a diverse group of photoreactives are available, diazirines are most frequently utilized because of their small size and high reactivity. Probes using diazirines are widely used to map protein interactions and reveal small molecule targets. In a practical context, such a technique can be used to screen medicinal drugs, natural products, and other kinds of small molecules.
Challenges in The Use of Diazirines-Based Probes
Despite their various advantageous qualities, diazirines still present significant challenges that limit their ability to be a preferred method for the determination of binding sites:
- Probe-labeled peptides tend to be low in number, which significantly decreases the probability of them being detected and analyzed.
- Cofragmentation of peptide backbones and probe adducts results in changes to the adduct’s mass and creates additional fragment ions, which makes it more difficult to match it with in silico predicted spectra.
- The false discovery rate (referring to the predicted portion of peptides that are assigned incorrectly) differs from unmodified peptides.
Finally, the insertion of reactive intermediates means that all locations in a peptide sequence are taken into account for adduction, thus resulting in a substantial increase in search space, elongated analysis times, decreased numbers of identifications, and an increased probability of producing false positives.
In order to enhance the utility of these probes, the spectral analysis of probe-labeled peptides requires improvement. It was found that the peptides tend to generate chimeric spectra as a result of the coisolation and coelution of different peptides that had the same sequences but were labeled on different amino acid residues. Probability models were constructed using this information to accurately predict whether a given probe-labeled peptide corresponds to a spectrum. The models were utilized to offer better coverage and location confidence.
The workflow also facilitates the construction of site-specific and proteome-wide concentration profiles for the probes in living cells, which were then confirmed using orthogonal techniques. This data was used to predict, and experimental structures were used in order to map thousands of binding pockets across the proteome, of which several had no reported ligands. The information garnered was then integrated with molecular docking for the generation of predictive binding modes for more than 80 interactions, thus aptly depicting the method’s usefulness in the realm of ligand design.
Diazirines-based analyses were first benchmarked utilizing conventional workflows. It was found that though some proteins interacted with various probes, the vast majority preferred to bind with a single probe or a single subset of them, thus indicating that fragment groups imparted binding preferences.
Over the course of the study, it was noticed that the spectra of labeled peptides had characteristics that differed from those of unlabeled peptides – it is possible that the spectra assigned to labeled peptides possess multiple peptides and are hence considered chimeric. Given their abundance, a predictive model was generated to predict the probability of a given spectrum containing a labeled peptide. The spectral features of probe-labeled and unlabeled peptides were compared systematically. The method is compatible with other proteome search engines, showing that the technique can be incorporated easily into many analysis pipelines.
A new analysis pipeline, Dizco (diazirine probe-labeled peptide discoverer), was then developed for the identification of probe-labeled sites. Its workflow uses information obtained from the chimeric spectra and utilizes predictive modeling in order to generate objective confidence measures for labeled peptides. The workflow was shown to yield a more comprehensive depiction of label locations in comparison to conventional analyses.
The increased confidence enabled by the use of Dizco makes the use of isotopically encoded enrichment tags unnecessary. Hence, the workflow was integrated with TMT-based quantitative proteomics in order to enhance quantitation, labeled peptide detection, and throughput across samples. The model was also shown to provide similar results when applied to different data, indicating its potential for use in a large range of experiments once properly trained.
Conclusion
Photoaffinity probe-labeled peptides are of great significance in the field of ligand binding, and the use of computational biology enables the creation of predictive models that utilize information on the probes’ characteristics to enhance the identification of binding sites as well as confidence. Significantly, it was discovered that these peptides generate chimeric spectra in MS analysis. Though the effects of the spectra can complicate the results of the experiments, accurately identifying them is a challenging task. The creation of a multiplexed, automated workflow – Dizco – facilitated the efficient construction of probe-protein concentration profiles within live cells. Experimental data, used in tandem with docking and structural information, can thus reveal the details of molecular characterization at various binding sites in the proteome.
Article Source: Reference Paper | Reference Article | Scripts developed in the study are available on GitHub and Zonodo
Learn More:
Sonal Keni is a consulting scientific writing intern at CBIRT. She is pursuing a BTech in Biotechnology from the Manipal Institute of Technology. Her academic journey has been driven by a profound fascination for the intricate world of biology, and she is particularly drawn to computational biology and oncology. She also enjoys reading and painting in her free time.