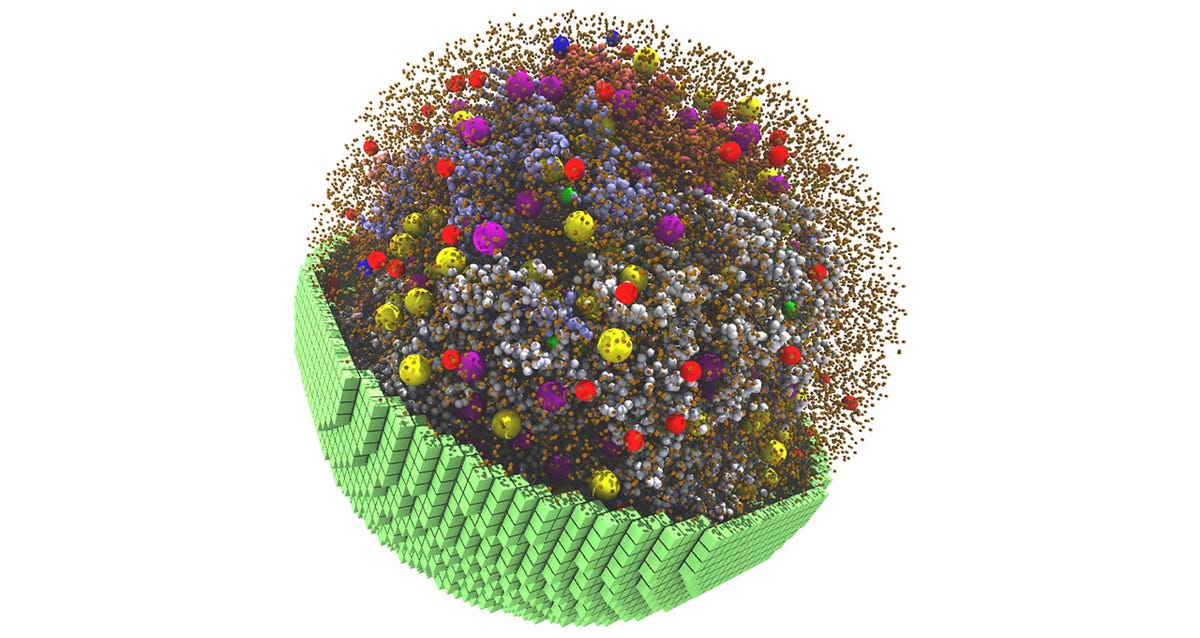
GPU-accelerated software developed by the researchers at the University of Illinois at Urbana-Champaign simulates a 2-billion-atom cell that metabolizes and grows like a biological cell. Researchers present a whole-cell fully dynamical kinetic model that replicates the physical and chemical characteristics at a particle scale of a minimal living cell-JCVI-syn3A.
The study of the simplest living cells may reveal principles that apply to all living systems, just like the study of hydrogen, the simplest atom, led to the knowledge of more complicated atoms. Consequently, the researchers are interested in investigating “minimum cells” by creating and constructing biological genomes devoid of non-essential genes in the lab and have been able to create living cells with fewer genes than any naturally occurring cell, making them easier to describe in detail.
Image Source: Fundamental behaviours emerge from simulations of a living minimal cell
The evolution of whole-cell models (WCMs), including how they evolved from genome-scale metabolic models (GSMMs) and how their steady-state fluxes were calculated, has been recently examined. The simulation of a living cell presented in this study, unlike earlier WCMs, is based on fully dynamical kinetic models in which subsystem networks and chemical species are continually interconnected over time on a single-cell basis.
An appropriate system for such a whole-cell model would be a minimal cell consisting of as few genes and reactions as possible for the cell to grow and divide. JCVIsyn3A is a genetically simple bacterial cell with just 493 genes on a single 543-kbp circular chromosome, 452 of which code for proteins, some of which are multi-domain complex components.
The scientists created a whole-cell kinetic model of Syn3A using a combination of stochastic and deterministic simulations using some background data. Ordinary differential equations (ODEs) are used to model the kinetics of the primary metabolic network, whereas stochastic simulations are used to model the kinetics of genetic information processing. They consider two stochastic simulation models: a chemical master equation (CME) description, which assumes that the entire cell is well stirred and implicitly includes the effects of diffusion in the rates of transcription, mRNA degradation, and translation; and a reaction-diffusion master equation (RDME) description, which requires macromolecules to diffuse to each other for reactions to take place in the cell’s spatially heterogeneous environment.
The researchers provide the results of the well-stirred model for 174 healthy replicating cells out of a total population of 207 cells during the course of a cell cycle. Meanwhile, the well-stirred model runs on CPUs and generates a whole-cell model that can be easily complexified to add additional pathways and testing.
The spatial model has a total of 7,765 distinct molecules and intermediates, as well as over 7,200 processes, such as RNAP binding to a gene start site. For this model, GPU-based Lattice Microbes software was used to simulate replicates over the first 20 minutes of a cell cycle that was stimulated for only eight cells. No significant growth or DNA replication was expected during this short time frame. The simulations reveal how many active degradosomes, RNAP, and ribosomes are there.
The spatial model has higher mRNA counts on average, likely due to the difference in transcription rates between the two models. The spatial model has lower nucleotide concentrations, which can be attributed to two factors: first, more nucleotides are being integrated into mRNA, and second, the current spatial model does not include DNA replication, which starts in the well-stirred model around 10 minutes on average. The somewhat higher counts of scaled proteins produced in the well-stirred simulations indicate incomplete replication events occurred in a few cells during the first 20 minutes. The active degradosome statistics in the spatial model determine the half-lives of well-stirred mRNA. The half-lives in the well-stirred model are longer than those in the spatial model, with average half-lives of 3.4 minutes and 1.97 minutes, respectively.
Simulations using the hybrid well stirred (CME-ODE) whole-cell kinetic model have already provided quantitative insight into how the cell balances the needs of metabolism, genetic information, and growth throughout a cell cycle. We may begin to comprehend the principles of life for this minimum cell when little regulation is presently based on the emergent behaviors coming from the well-stirred and spatially resolved stochastic-deterministic simulations shown here.
To calculate the behavior of a cell over a maximum cell cycle of 120 minutes, the hybrid CME-ODE whole-cell kinetic model takes about 4 hours. Before moving on to the full spatial hybrid RDME-CME-ODE model, which is computationally more demanding, the hybrid model can be easily complicated by adding additional reactions.
The explicit tracking of every energy molecule utilized by active reactions in metabolic and genetic information processing is crucial for this approach. The researchers used fully dynamical kinetic modeling to enhance the ATP cost calculations for the cellular networks of Syn3A (containing both metabolism and genetic information processing) to single-cell and single-reaction resolution as a function of time. At each time step, the total ATP generated is close to or slightly larger than the total ATP used. They know their exact ATP costs by tracking the fluxes through each reaction because each transport reaction is simulated independently. Syn3A has been reduced to the point that it must transport its building blocks, unlike most organisms that have synthesis pathways for most of their building blocks. Metabolic processes have the highest cost in Syn3A, with the Phosphofructokinase reaction in upper glycolysis accounting for 75% of the overall metabolic cost.
In the future, the researchers plan to investigate numerous cell cycles that begin with daughter cells, each possibly containing a chromosome with multiple replication forks. They could get statistics about cell divisions and multiple initiations of DNA replication events and DNA filament formation by extending the simulations over several cell cycles. Regulation is also crucial; therefore, future models will contain it. The scientists will modify the spatially resolved kinetic model to allow for changes in cell shape, DNA replication, and ribosome biogenesis, and these processes will be addressed.
Story Source : Thornburg, Z. R., Bianchi, D. M., Brier, T. A., Gilbert, B. R., Earnest, T. M., Melo, M. C., … & Luthey-Schulten, Z. (2022). Fundamental behaviors emerge from simulations of a living minimal cell. Cell, 185(2), 345-360.
Data and code availability: https://doi.org/10.1016/j.cell.2021.12.025
Background Image: Snapshot from the 20-minute 3D spatial simulation, showing yellow and purple ribosomes, red and blue degradasomes, and smaller spheres representing DNA polymers and proteins.
Background Image Source:https://blogs.nvidia.com/blog/2022/01/20/living-cell-simulation/
Learn More:
Top Bioinformatics Books ↗
Learn more to get deeper insights into the field of bioinformatics.
Top Free Online Bioinformatics Courses ↗
Freely available courses to learn each and every aspect of bioinformatics.
Latest Bioinformatics Breakthroughs ↗
Stay updated with the latest discoveries in the field of bioinformatics.
Swati Yadav is a consulting intern at CBIRT. She is currently pursuing her Master's from Delhi Technological University (DTU), Delhi. She is a technology enthusiast and has a keen interest in the latest research on bioinformatics and biotechnology. She is passionate about exploring new advancements in biosciences and their real-life application.