Researchers from Rice and Boston universities developed CLASSIC, a platform that combines long- and short-read sequencing to measure over 100,000 multi-gene circuits in human cells. The resulting data was used to train machine learning models to predict circuit behavior and uncover rules for part compositions, accelerating the design-build-test-learn cycle in synthetic biology.
Limitations of the Traditional Synthetic Biology Workflow
Synthetic biology aims to engineer living cells with new predictable functions by assembling genetic parts into complex gene circuits. A problem arises because genetic parts interact in complex context-dependent ways inside crowded cells; circuit performance is difficult to predict, often requiring many iterative design-build-test-learn (DBTL) cycles. To support this process, researchers use high-throughput screening approaches that test large libraries of gene circuits simultaneously and generate data.
However, most existing high-throughput methods rely on short read sequencing, which works well for short DNA elements but becomes costly and technically challenging for long DNA constructs that encode complete gene circuits. Barcode allows short-read sequencing to measure the activity of long DNA constructs, but it still requires complex and low-throughput workflows to link each barcode to its corresponding construct, limiting scalability and flexibility. Even though all these strategies exist, they have only partially addressed the issue and are limited by complex assembly workflow, reduced library design flexibility, reports, and practical constraints on construct length and library size.
What is CLASSIC?
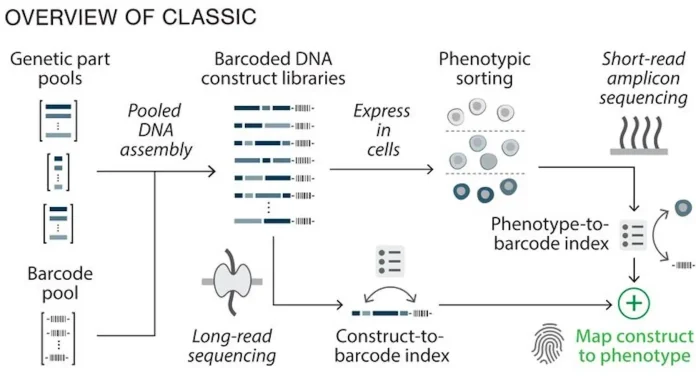
CLASSIC is a novel high-throughput genetic screening platform that combines long-read Nanopore sequencing with short-read Illumina sequencing to analyze complex multi-gene DNA constructs in living cells. Nanopore sequencing is used to decode the full genetic composition of each construct. In contrast, Illumina sequencing is used to measure functional performance, enabling ultra-high throughput and quantitative mapping of genotype-phenotype relationships. Traditional DBTL tests a few designs to learn rules, while CLASSIC tests thousands at once and lets data plus AI discover the rules.
Single-Input Circuit Library
Researchers constructed a large library of inducible two-gene circuits containing more than 100,000 variants. Each circuit consists of two expression units: one encoding a synthetic zinc finger transcription factor (synTF) and the other encoding a GFP reporter gene activated only when the synTF binds to specific DNA sites before the gene. The transcript factor is linked to a modified estrogen receptor that responds to the drug 4-hydroxytamoxifen (4-OHT). When the drug is added, the transcription factor enters the nucleus and turns on the reporter gene; without the drug, the reporter stays off.
The aim was to achieve high fold-change, low background, and strong induction, which is difficult in mammalian cells due to complex regulatory tuning. CLASSIC was used to quantify how different circuits affect this balance. Nanopore sequencing was used to determine the full composition of each construct and link barcodes to most circuit designs. The library was inserted into cells, which were then sorted by fluorescence in the presence and absence of 4-OHT. Sequencing the barcode with Illumina revealed the expression and changes for thousands of circuits, confirming that the CLASSIC method provides accurate measurements for large libraries.
Machine Learning Prediction
Because 27% of circuit designs were unmeasured, machine learning models were trained to predict their behavior. The multilayer perceptron performed best. This model was used to predict basal-induced expression and fold-change expression for all 165,888 designs, revealing that most circuits have low fold change, and only about 8% achieved high fold change overall. Prediction accuracy was low for extreme behaviors, including high fold-change variants. To validate the accuracy of the model predictions, cell lines containing both previously measured and unmeasured circuit designs were tested, and flow cytometry results closely matched the predictions, strongly confirming the validity of both groups and validating many high-fold change designs.
The model also corrected apparent measurement errors, showing that the CLASSIC-trained ML model can reliably predict the behavior of unmeasured or inaccurate limited gene circuits. The ML model’s SHAP analysis revealed that 6 component categories—activation domains,IDPs, ZF affinity, and promoters—are primarily responsible for high fold change (HFC) behavior. These components are highly interdependent, and clustering identifies 3 different high-fold-change designs that share locus features, minimizing basal expression while tuning transcription factor properties. Overall, HFC circuits achieve low background and strong induction through coordinated part selection and locus design.
Model Generalization and Multi-input Circuit Expansion
It was found that only 9% of the training data is sufficient to achieve near-maximum prediction accuracy; the model demonstrated strong generalization and scalability. To test its generalization, researchers designed gene circuits with two drug-controlled transcription factors regulating a single reporter gene, enabling AND/OR-like logic. The model was trained using data from a densely sampled base library and an expansion library that introduced additional features. The model’s predictions for circuit behavior were then experimentally tested by constructing and measuring selected predicted designs, and the close agreement confirmed the model’s accuracy and demonstrated that the CLASSIC ML framework can scale to extremely large and complex genetic design spaces.
Conclusion
CLASSIC enables high-throughput quantitative profiling of complex multi-kb genetic part combinations in cells. It greatly expands the scale of genetic design spaces that can be explored using time, cost, and DBTL cycles in synthetic biology projects. While traditional methods might be more economical for smaller projects, CLASSIC is best suited for medium-to large-scale or underdetermined design problems. The data generated by CLASSIC can be used to train machine learning models that generalize to much larger design spaces, paving the way for AI-driven and potentially generative gene circuit design across a range of biological systems.
Article Source: Reference Paper bioRxiv | Abstract | Code Availability: GitHub
Disclaimer:
The research discussed in this article was conducted and published by the authors of the referenced paper. CBIRT has no involvement in the research itself. This article is intended solely to raise awareness about recent developments and does not claim authorship or endorsement of the research.
Follow Us!
Learn More:
Jainab Shaikh is a postgraduate in Biotechnology with a strong interest in understanding how research translates into real-world innovation. Her areas of focus include biosensors, bioinformatics, and sustainable biotechnological applications. She is passionate about exploring recent scientific advancements and communicating them through clear, engaging, and accessible content. Her work particularly emphasizes research-driven narratives in healthcare, biotechnology, skincare science, and emerging life science innovations.