Microsoft researchers introduce BioEmu-1, an updated version of BioEmu that advances the knowledge of how proteins function by offering a window into the diverse range of structures that each protein can take on or structural ensembles. BioEmu is a generative deep learning system that tackles the scientific challenge of forecasting dynamical dynamics in proteins following structural revolutions. Using vast volumes of data and cutting-edge training methods, this system produces thousands of statistically unique protein structure ensemble samples every hour. Protein conformations are quantified by BioEmu with relative free energy errors of roughly 1 kcal/mol, yielding empirically testable hypotheses and mechanistic insights like the causes of fold instability. This significant area of research is essential to comprehending the dynamics and architectures of proteins. Many treatments work by altering protein structures to improve their function or stop them from harming people. Thus, a deeper understanding of proteins helps us create more effective medications.
Introduction
In humans and other living creatures, proteins are vital for nearly every biological function, from building muscle fibers to preventing illness. The ability to accurately predict protein structures from their amino acid sequences has been made possible by the remarkable advancements made in recent years in deep learning for a better knowledge of protein structures. However, looking at a single frame of a movie is similar to anticipating a single protein structure based on its amino acid sequence; it merely provides a glimpse of a very flexible molecule.
Molecular dynamics (MD) simulations are one method of modeling various protein structures. These methods, which are widely used in both academia and industry, mimic the movement and deformation of proteins throughout time. However, MD simulations need to run for a long time to replicate structural changes that are functionally significant. This process requires a lot of calculation, and a lot of work has been done to speed up simulations, even going so far as to create unique computer architectures. However, despite these advancements, many proteins are still beyond present simulation capabilities and would take years or even decades to complete.
BioEmu-1
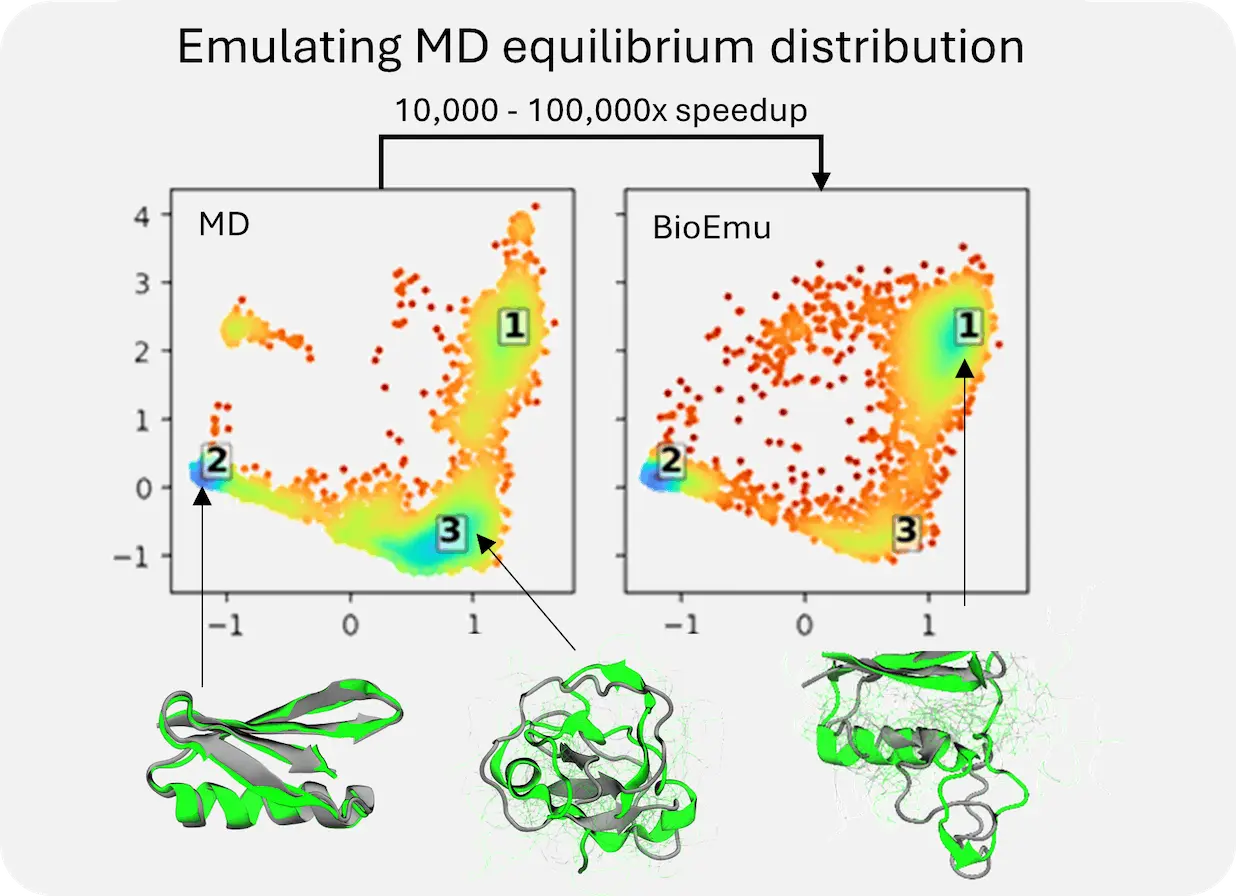
A single graphics processing unit can produce hundreds of protein structures every hour using the deep learning model BioEmu-1. Researchers are making BioEmu-1 an open-source like the previous BioEmu model to enable protein scientists to use the model to analyze structural ensembles. BioEmu, also known as the biomolecular emulator, is a novel machine learning system that can accurately sample from the equilibrium distribution of protein conformations in a few GPU hours of each experiment. It offers orders of magnitude more computing efficiency than traditional MD simulations, which makes it possible to gain insights that were previously unattainable. BioEmu-1 is part of Azure AI Foundry Labs, a platform where developers, startups, and businesses may investigate cutting-edge discoveries from Microsoft research.
This has been made possible by using three different kinds of data sets to train BioEmu-1: (1) AlphaFold Database (AFDB) structures, (2) a large dataset of MD simulations, and (3) a dataset of experimental protein folding stability. Related protein sequences are clustered throughout the training phase to identify unique structures. The MD simulation dataset aids in predicting structural changes surrounding these islands to map out the several alternative configurations a single protein can adopt. BioEmu-1 is adjusted to sample folded and unfolded structures with the appropriate probabilities using the protein folding stability dataset.
Together, these developments enable BioEmu-1 to predict many structures and generalize to unknown protein sequences. The LapD protein from the Vibrio cholerae bacteria, which causes cholera, can have its structures predicted by BioEmu-1. Both the bound and unbound structures of LapD with c-di-GMP molecules, which are not present in the training set but are known experimentally, are predicted by BioEmu-1. The model also provides a perspective on intermediate structures that have never been detected experimentally, leading to plausible theories regarding the function of this protein. Further developments in fields like drug development are made possible by understanding how proteins function.
Conclusion
Protein stability, which scientists evaluate by calculating the folding free energies—a method of expressing the ratio between a protein’s unfolded and folded states—is precisely predicted by BioEmu-1. When developing proteins, for example, for therapeutic applications, protein stability is a crucial consideration. Researchers observe that the projected free energy values show good agreement with experimental values, even on sequences that BioEmu-1 has never encountered during training. By using generative deep learning to enable lightning-fast sampling of the free-energy landscape of proteins, BioEmu represents a major advancement in this direction. Scientists think that BioEmu-1 is a starting step in creating the entire set of possible protein structures. Researchers are also conscious of its limits in these early stages. Scientists anticipate that by making BioEmu-1 open-source, they will begin to experiment with it and help us identify its strengths and weaknesses so that researchers can make improvements in the future.
Article Source: Reference Article | GitHub.
Disclaimer:
The research discussed in this article was conducted and published by the authors of the referenced paper. CBIRT has no involvement in the research itself. This article is intended solely to raise awareness about recent developments and does not claim authorship or endorsement of the research.
Follow Us!
Learn More:
Deotima is a consulting scientific content writing intern at CBIRT. Currently she's pursuing Master's in Bioinformatics at Maulana Abul Kalam Azad University of Technology. As an emerging scientific writer, she is eager to apply her expertise in making intricate scientific concepts comprehensible to individuals from diverse backgrounds. Deotima harbors a particular passion for Structural Bioinformatics and Molecular Dynamics.