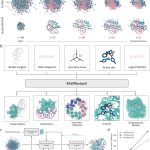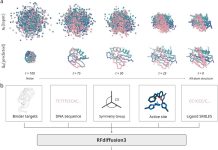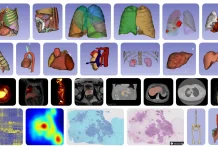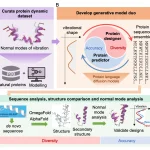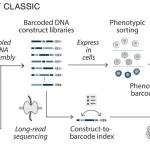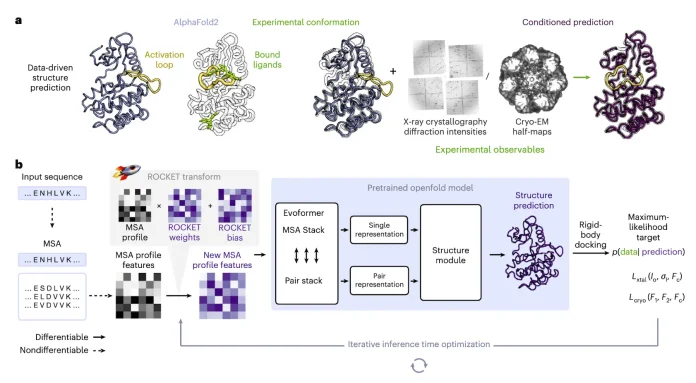
Researchers at the University of Cambridge, Columbia University, and partner institutions have introduced ROCKET, a novel framework that extends AlphaFold2 by integrating experimental data, such as cryo-EM and x-ray crystallography, directly into structure prediction. It fuses machine learning priors with traditional approaches, allowing accurate modeling even at low resolution, noisy, or frontier datasets. Rocket was published in Nature Methods in the first week of April.
Models like AlphaFold2, RoseTTAFold, IntelliFold-2, and similar models were trained on decades of painstakingly collected experimental structures obtained through X-ray crystallography, cryo-EM, and NMR. But these models are only as good as the structural data they were trained on. As high-quality datasets capturing multiple conformations and interactions are scarce, these models predict protein structures with near-experimental accuracy but still struggle with side-chain packing, functional dynamics of proteins, conformational flexibility, and noisy low-resolution data to this day.
Understanding the Bottleneck Addressed by ROCKET
Experimental data often come in the form of noisy maps or diffraction patterns. To make sense of these datasets, scientists must reconstruct atomic models, which is a slow and error-prone process, especially when the resolution is low.
Current workflow involves a starting structure from ML predictions like AF2, which are refined by software like phenix. refine to fit experimental data, along with pattern-matching tools that help correct local mismatches. But this OTC solution breaks down when it comes to large rearrangements in the protein (secondary structure switches, domain shifts, etc.) or when the fine details blur out (at resolutions < 4-5 Å), making it hard even for expert human modelers.
If the machine learning priors like those embedded in AlphaFold can be integrated directly into refinement, they could:
- Guide model building in noisy/low-resolution datasets
- Reduce the need for manual intervention
- Yield more accurate atomic models in frotien cases.
This was exactly the motivation behind refining OpenFold with Crystallographic/cryo-EM likelihood targets. The team highlights that ML has revolutionized the field, but it’s not a replacement for experiments. The future lies in integrating both, which is exactly what ROCKET does by conditioning AlphaFold predictions on experimental observables.
Core Innovation Behind ROCKET: Existing Attempts and ROCKET’s Approach
The idea of incorporating structural priors into ML isn’t new. Previous attempts, like ColabDock’s incorporation of crosslinking restraints into complex predictions, worked well in some cases but weren’t as robust for challenging tasks.
Adjusting AlphaFold’s weights to incorporate experimental data opened the door to new challenges. Retraining is computationally expensive, demands large datasets, and is limited to a specific modality, for example, cryo-EM only, and training new architectures like ModelAngelo was not generalizable either. So researchers came up with a ‘Predict And Build’ approach.
ROCKET, however, combines OpenFold, a reimplementation of AF2. It is different because it doesn’t retrain or decouple steps. Instead, it performs inference-time optimization directly in AF’s latent space, keeping predictions and refinements tightly coupled and guided by likelihood functions. AF2 can be expanded to explore different conformations of a protein. ROCKET borrows this data and optimizes the embedded multiple-sequence alignment (MSA) cluster profile. By tweaking it, ROCKET can move predictions toward experimentally supported conformations smoothly.
Simons Foundation describes ROCKET as a major upgrade to AlphaFold, allowing it to “learn from raw experimental data” and tackle protein folding challenges linked to disease like cancer.
Impact: Why this Research is Valuable
- High Resolution Benchmarks: When tested on 27 crystallographic structures, ROCKET solved AF’s cutoff. It consistently improved AF’s prediction, reducing backbone shifts and improving side-chain fit.
- Low-Resolution Cryo-EM tests: For the SLC19A3 thiamine transporter, it recovered the correct outward open conformation even at 10Å resolution, where side chains are fuzzy. For complexes like GroEL: GroES-ATP, it managed to capture large domain rearrangements at 2.9 Å and 4.9 Å, but it struggled at 6.8 Å. But most importantly, it sometimes outperformed manual expert refinements, proving its strength.
- Case Studies: Cryo-ET averages, low-resolution crystallography, prefers particle orientations. In each case, ROCKET produced models that aligned with experimental data.
Key Takeaways
AlphaFold2 predicts protein structure with absolute accuracy but finds it hard to work with side-chain packing and noisy and low-resolution data. Refining OpenFold with Crystallographic/Cryo-EM likelihood targets (ROCKET) incorporates experimental data directly into AF’s inference, breaking the low-resolution barrier with no retraining required. It’s a step forward in closing the loop between computation and experiments.
Article Source: Reference Paper
Disclaimer:
The research discussed in this article was conducted and published by the authors of the referenced paper. CBIRT has no involvement in the research itself. This article is intended solely to raise awareness about recent developments and does not claim authorship or endorsement of the research.
Follow Us!
Learn More:
Saniya is a graduating Chemistry student at Amity University Mumbai with a strong interest in computational chemistry, cheminformatics, and AI/ML applications in healthcare. She aspires to pursue a career as a researcher, computational chemist, or AI/ML engineer. Through her writing, she aims to make complex scientific concepts accessible to a broad audience and support informed decision-making in healthcare.