Researchers from the Broad Institute of MIT and Harvard, Harvard University, Carnegie Mellon University, and Howard Hughes Medical Institute, the study introduces Lyra—a groundbreaking deep-learning model designed for efficient biological sequence modeling.
The Need for Efficient Biological Sequence Modeling
Understanding the intricate relationships between DNA, RNA, and protein sequences is at the heart of modern computational biology. Since the evolution of biology-based algorithms, deep learning models like Convolutional Neural Networks (CNNs) and Transformers have emerged capable of analyzing biological sequences. CNNs usually capture local patterns within sequences, while the Transformer’s attention mechanism is more appropriate for long-range dependencies. Nevertheless, the majority of biological applications cannot afford the necessary resources because of the extreme computational costs associated with models based on Transformers.
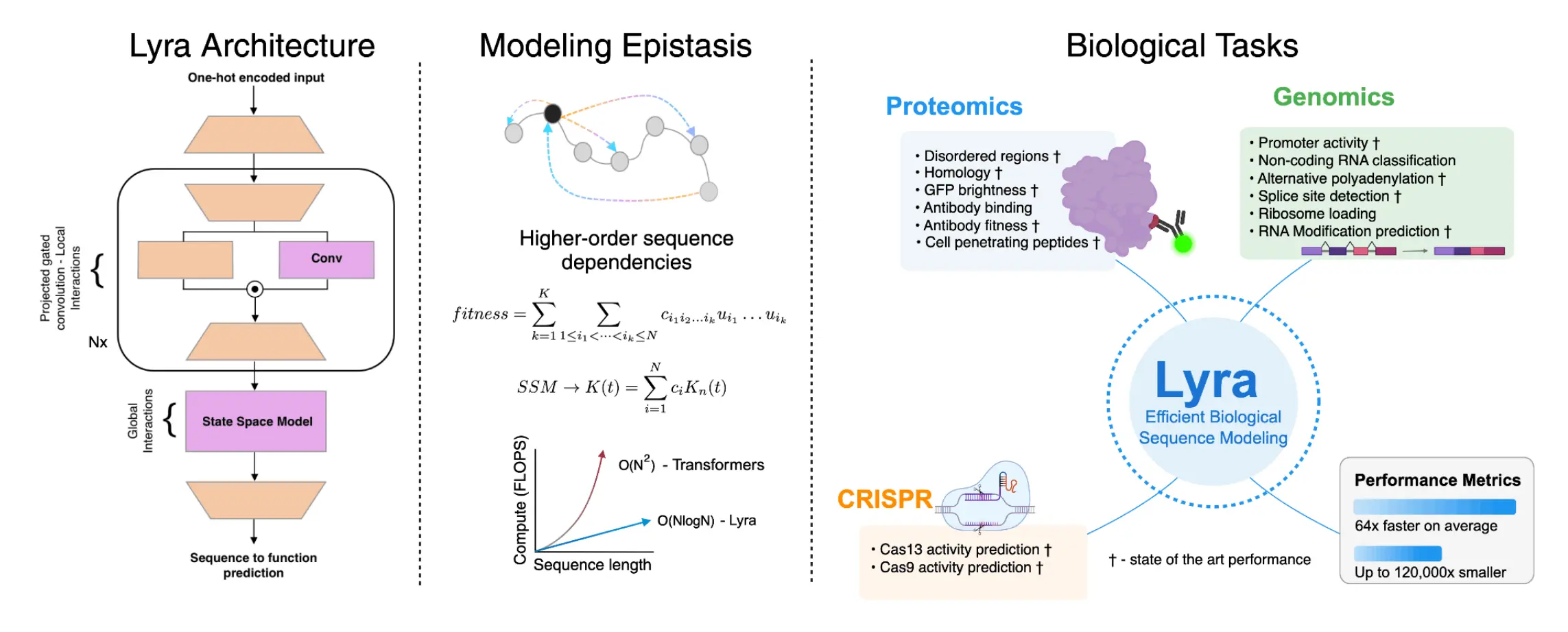
There is a need for new models that operate within a reasonable computational budget but, at the same time, do not trade off efficiency. This is what Lyra seeks to solve; Lyra is an artificial intelligence model that incorporates state space models (SSMs) with projected gated convolutions (PGCs) to sufficiently take into account both local and global interactions within the sequence.
What Makes Lyra Different?
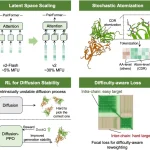
Lyra’s architecture is designed to align with the principles of biological sequence interactions, particularly epistasis—the influence of genetic mutations on each other. Unlike transformer models that are self-referential and have quadratic complexity, Lyra utilizes SSMs, which are more complex but scale more efficiently with sequence length. Lyra incorporates PGCs to ensure a thorough capture of local sequence features along with long-range sequence comprehension via SSMs.
Lyra embodies epistasis by utilizing a polynomial-based framework for complex sequence interactions, making it energy-efficient for a wide span of biological tasks. This approach simultaneously alleviates the processing requirements while enhancing the understanding and answerability of the sequence-function relationships.
Lyra’s Performance Across Biological Tasks
The research team tested Lyra across more than 100 biological tasks, demonstrating its superior performance in key areas such as:
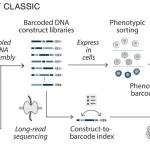
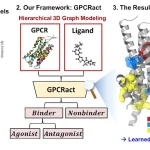
- Protein Fitness Prediction: Lyra accurately predicts how mutations affect protein function, outperforming Transformer-based models in tasks like antibody binding and GFP fluorescence prediction.
- RNA Structure and Function Analysis: From predicting RNA secondary structures to identifying functional elements like splice sites, Lyra showcases state-of-the-art (SOTA) accuracy while using significantly fewer computational resources.
- CRISPR Guide Design: The model enhances genome editing by improving Cas9 and Cas13 guide RNA predictions, ensuring higher precision in genetic modifications.
- Peptide Engineering: Lyra is effective in designing peptides with desired biophysical properties, such as cell-penetrating peptides crucial for drug delivery.
The most notable trait of Lyra is her efficiency in computing. She does not need tons of data and clusters of GPUs to be trained, as she outshines other large transformer models. For example, Lyra achieves SOTA results with a considerably smaller number of parameters, which makes her outperform ProtT5 and ESM-1v models that require up to 120,000 times more parameters.
Breaking Computational Barriers
Traditional deep-learning biological sequence models are tied to heavy hardware dependencies, which only a few well-funded labs can afford. In contrast, Lyra trains and performs inference on sequence models in a few hours using only two consumer-grade graphic processing units, bringing advanced sequence modeling within reach of hundreds of researchers. This is revolutionary in enabling rich biological sequence modeling for users globally.
Additionally, Lyra’s computational efficiency allows for the processing of much longer sequences—up to 65,536 base pairs—something that Transformer models struggle with due to their quadratic complexity. This capability is crucial for studying long genomic sequences and regulatory elements that govern gene expression.
Conclusion
Lyra leads the way in computational biology, enabling rich, scalable, and easy functionality for advanced biological sequence modeling in comparison with the other transformer models. sAI continues to evolve the life sciences. Lyra, with its principle mathematically based architecture, aids in tackling the issues concerning biological sequence and stands firm to the expectations.
Article Source: Reference Paper.
Disclaimer:
The research discussed in this article was conducted and published by the authors of the referenced paper. CBIRT has no involvement in the research itself. This article is intended solely to raise awareness about recent developments and does not claim authorship or endorsement of the research.
Important Note: arXiv releases preprints that have not yet undergone peer review. As a result, it is important to note that these papers should not be considered conclusive evidence, nor should they be used to direct clinical practice or influence health-related behavior. It is also important to understand that the information presented in these papers is not yet considered established or confirmed.
Follow Us!
Learn More:
Anchal is a consulting scientific writing intern at CBIRT with a passion for bioinformatics and its miracles. She is pursuing an MTech in Bioinformatics from Delhi Technological University, Delhi. Through engaging prose, she invites readers to explore the captivating world of bioinformatics, showcasing its groundbreaking contributions to understanding the mysteries of life. Besides science, she enjoys reading and painting.