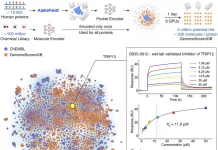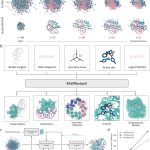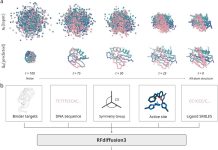
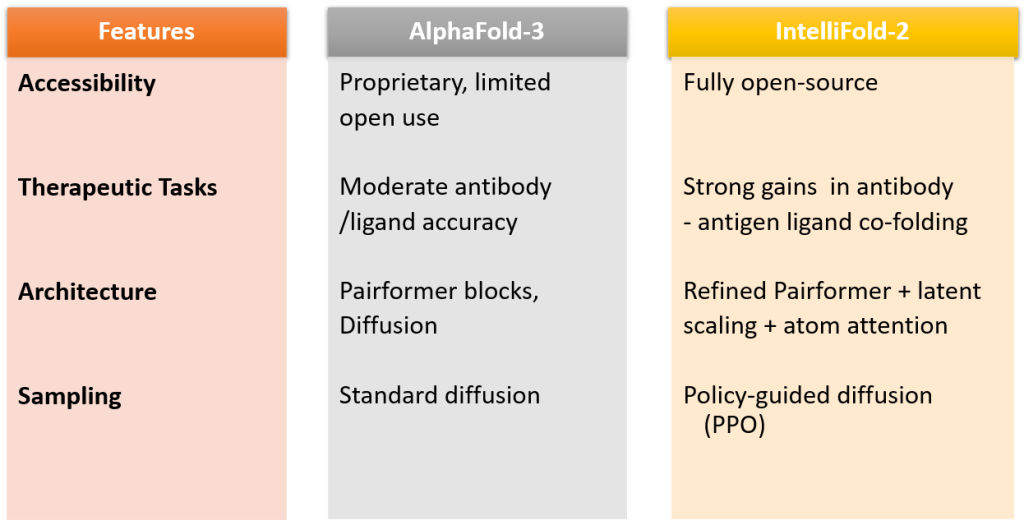
The innovators behind intelliFold-2 at IntelliGen AI emphasize that IntelliFold-2 is designed as a state-of-the-art open source alternative to AlphaFold3 with stronger performance in therapeutically relevant tasks like antibody-antigen docking and protein-ligand co-folding. The architectural refinements, like latent scaling and atom attention, along with algorithmic innovations such as policy-guided diffusion and difficulty-aware loss reweighting, help it directly target AlphaFold3’s weak spots, making it more robust.
Biomolecular structures like proteins, RNA, and complexes are the blueprints of life at the molecular level. They tell us how a protein’s 3D shape binds to other molecules, what interactions happen at the atomic level, and how these reactions can be catalysed, etc., which is essential for scientists to develop new and effective therapeutics.
Without accurate structures, biology becomes guesswork. We can understand the sequence of a protein, but not how it folds or interacts, which limits our ability to design drugs and vaccines and to understand disease pathways.
The Need for Accurate Biomolecular Structures: Why AlphaFold-3 Was Not Enough
Older methods like X-ray crystallography and NMR were slow, expensive, and limited. It took years for scientists to solve one structure using these methods. AI models like AlphaFold-3 were revolutionary because they expanded the scope to handle complexes, ligands, and interactions with near experimental accuracy.
But biology isn’t just about lone proteins floating around. Most of the actions happen when proteins interact with other proteins, small molecules, antibodies, RNA, or membranes. These interactions are much harder to predict because they involve flexible shapes, chemical environments, and dynamic bindings.
Developers of IntelliFold-2 highlight two particular challenges as the motivating problems the model addresses. These are:
- Antibody-antigen Interactions: These interactions are central to vaccines and immunotherapies, but prediction here is the hardest yet most useful.
- Protein-Ligand Co-Folding: This is the foundation of drug discovery. However, the dynamic nature of proteins makes it much harder to predict their structures.
These are the hardest problems in biomolecular prediction, and they are also the ones directly tied to medicine. IntelliFold-2 deliberately targets these weak spots with architectural refinements and smarter sampling.
Siqi Sun, the tech lead, shared on LinkedIn that IntelliFold-2 was built to outperform AlphaFold-3 on critical therapeutic tasks. Unlike AlphaFold-3, Intellifold-2 is fully available for academic use with three variants (Flash, v2, Pro).
Why IntelliFold-2 Handles The Challenges Better: Some Core Technical Innovations in IntelliFold-2
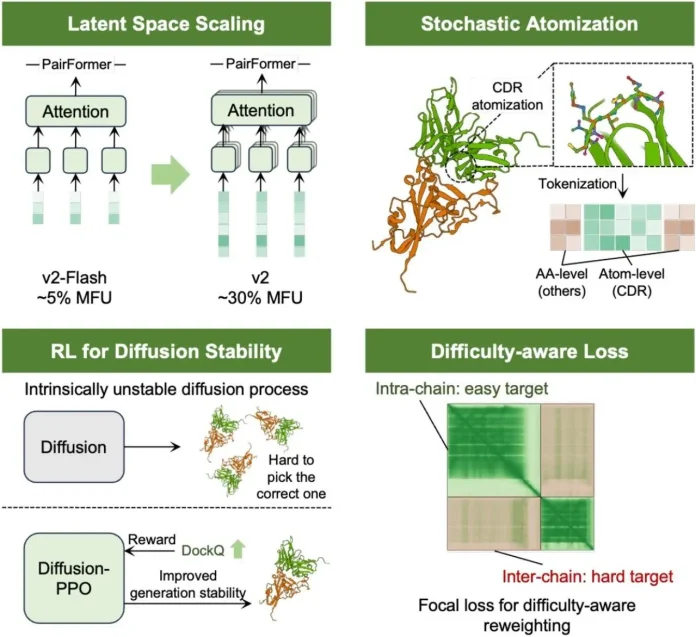
Latent Space Scaling in Pairformer blocks: The latent space within Pairformer blocks encodes relationships between atoms and residues, but was too narrow to handle the structural details the model could represent, and also wasted GPU efficiency. By increasing the dimensionality of these latent representations, IntelliFold-2 can capture more complex biological interactions.
Principled Atom Attention and Stochastic Atomization: Proteins and biomolecules can be described at different levels, from individual atoms to whole chains. The revised ‘atom attention’ mechanism ensures that predictions at the atomic level don’t contradict the overall 3D structure. Also, instead of tokenizing the atoms in the same way, the model introduces randomness during training. This forces the model to learn more vigorous patterns.
Policy-Guided Sampling: In structure prediction, diffusion models generate conformations step by step, gradually refining them; here, the authors treat the diffusion sampler as a policy in reinforcement learning. This means each sampling trajectory is like a decision path.
Foldbench Benchmarks: IntelliFold-2 V/S AlphaFold-3
IntelliFold-2 has proven itself against several categories of biomolecular structure prediction tasks and introduced itself as the next step beyond alphaFold-3:
- Protein Monomers (single): IntelliFold-2 slightly improves accuracy (0.89 vs 0.88 LDDT), showing it can capture finer details of individual proteins.
- RNA Monomers: IntelliFold-1 outperforms AlphaFold-3, but the flash version underperforms. RNA still remains a tough category.
- Protein-RNA interactions: IntelliFold-2 shows a clear gain over AlphaFold-3 (68.3% vs 62.3%), important for understanding gene regulation and viral replication.
Conclusions
AlphaFold-3 was a breakthrough, but there’s still room for improvement in challenging therapeutic contexts. IntelliFold-2 is positioned as a major step forward for the open-source community. The model is bigger and more expressive. It edges out AlphaFold 3 in monomer accuracy and protein-RNA interactions, while maintaining competitive performance in other categories. Its strongest gains are in antibody-antigen docking and protein ligand binding, two of the most medically relevant tasks.
Article Source: Reference Paper | Code Availability: GitHub | IntelliFold-2-Pro is available to Pro users at: Server
Disclaimer:
The research discussed in this article was conducted and published by the authors of the referenced paper. CBIRT has no involvement in the research itself. This article is intended solely to raise awareness about recent developments and does not claim authorship or endorsement of the research.
Important Note: bioRxiv releases preprints that have not yet undergone peer review. As a result, it is important to note that these papers should not be considered conclusive evidence, nor should they be used to direct clinical practice or influence health-related behavior. It is also important to understand that the information presented in these papers is not yet considered established or confirmed.
Follow Us!
Learn More:
Saniya is a graduating Chemistry student at Amity University Mumbai with a strong interest in computational chemistry, cheminformatics, and AI/ML applications in healthcare. She aspires to pursue a career as a researcher, computational chemist, or AI/ML engineer. Through her writing, she aims to make complex scientific concepts accessible to a broad audience and support informed decision-making in healthcare.